Main
Globally, CRC ranks as the third leading cancer type and the second most common cause of cancer-related death1. As CRC is an aging-associated disease, the rates of CRC incidence and death grow steadily with age, with an estimated 90% of worldwide cases and deaths occurring in people over 50 years old2. However, in recent years, worldwide cancer registries have reported a disproportionate increase in the incidence of EOCRC, generally defined as CRC diagnosed in people younger than 50 years of age3,4
EOCRC presents unique clinical and pathological characteristics in comparison with CRC in older patients, with a predominance of rectal and left colon tumors, higher prevalence of synchronous and metachronous CRC, higher frequency of metastatic disease at diagnosis and more aggressive and less differentiated tumors5. However, genomic alterations are largely similar to those described in older patients, although several genes exhibit different alteration rates6.
As our understanding of risk factors for CRC expands, so does the evidence that lifestyle factors, such as diet and exercise, can substantially influence CRC risk. This has led to the hypothesis that changes in lifestyle and environmental exposures—collectively known as the exposome—may play a critical role in the rising incidence of CRC in young patients. Nonetheless, efforts to identify modifiable risk factors specific to EOCRC have met with only limited success7, as most studies compare EOCRC cases with controls and identify the same risk factors as those for LOCRC cases8. This limited success could be attributed to the lack of quantitative measurements of exposome traits in cancer cohorts, which consequently makes it challenging to establish clear associations with cancer risk. Emerging studies have identified exposome-induced changes in CpG site-specific DNA methylation levels9, providing a new avenue for investigation. As DNA methylation data is frequently available in cancer cohorts, exposome-related methylation changes could serve as a valuable proxy for direct exposome measurements. This approach may enhance our ability to pinpoint specific exposome risk factors and improve our understanding of their role in EOCRC development.
In this study we aimed to explore the exposome traits that may contribute to the development of EOCRC compared to LOCRC, herein defined as CRC diagnosed in people 70 years and older. To this end, we constructed methylation risk scores (MRSs) for lifestyle and environmental factors using DNA methylation data from The Cancer Genome Atlas (TCGA) as the discovery cohort and conducted a replication meta-analysis across nine independent cohorts. The resulting MRSs were then compared between EOCRC and LOCRC. The relationship between pesticide use intensity and EOCRC incidence in the USA was further investigated using population-based data.
Results
Cohort description
For the discovery phase of this study, we utilized colon adenocarcinoma (COAD) samples obtained from TCGA10. The subsequent replication phase was carried out through a meta-analysis, incorporating data from studies on colon cancer (GSE131013, GSE42752, E-MTAB-7036 and GSE199057)11,12,13,14, rectal cancer (TCGA-READ and GSE39958)15 and colorectal cancer (GSE101764, GSE77954 and E-MTAB-3027)16,17,18. Patients were classified as EOCRC (<50 years) or LOCRC (≥70 years). This resulted in a discovery dataset containing 31 EOCRC and 100 LOCRC patients, whereas the replication cohort incorporated 83 EOCRC and 272 LOCRC patients. Population characteristics are presented in Table 1, with a detailed discovery cohort characterization in Supplementary Table 1.
Exposome-related DNA methylation marker sets
The exposome’s association with EOCRC versus LOCRC was evaluated using 29 lifestyle and environmental factors, which we define here as the exposome, acknowledging its broader scope. The analyzed traits encompassed 11 lifestyle factors: the Alternative Healthy Eating Index (AHEI), alcohol consumption, birthweight, body mass index (BMI) (continuous variable in kg m−2), cannabis use, coffee consumption, education level, Mediterranean Diet Score (MDS), obesity (defined as ≥30 kg m−2), smoking habits and smoking inference model (smoking-Maas). Furthermore, we examined four air pollution particles: nitrogen dioxide (NO2), polychlorinated biphenyls (PCBs) and particulate matter (PM) < 2.5 µm in diameter (PM2.5) and between 2.5 µm and 10 µm (PM2.5–10). In addition, we included 14 pesticides encompassing 2,4-dichlorophenoxyacetic acid (2,4-D), atrazine, acetochlor, chlordane, dicamba, malathion, dichlorodiphenyltrichloroethane (DDT), heptachlor, lindane, glyphosate, mesotrione, metolachlor, picloram and toxaphene. For the marker selection, we selected for each trait significantly associated CpG sites (CpGs) from extensive epigenome-wide association studies (EWAS), employing various significance thresholds, namely genome-wide (GW: P < 1.2 × 10−7), P < 1.0 × 10−5 and false discovery rates (FDR) of <0.01, <0.05 and <0.1. Smoking-Maas represents the CpGs included in a previously developed smoking inference model and their published coefficients19; no additional selection threshold was applied in the current study.
Exposome-related MRSs
Using available EWAS summary statistics across the five marker selection thresholds, we identified 63 exposome CpG sets across 29 exposome traits and computed 63 weighted MRSs from DNA methylation beta-values adjusted for Horvath epigenetic age20. Although termed ‘risk scores,’ these MRS serve as proxies for exposome exposure, enabling the assessment of exposure prevalence in early- versus late-onset tumors rather than estimation of individual cancer risk. The number of CpGs and their weights can be found in Supplementary Tables 2 and 3.
The limited availability of exposome data in tumor samples hinders the evaluation of its impact on EOCRC and restricts direct validation of the constructed MRSs against measured exposures. To address this, we performed a comprehensive validation of the constructed MRSs across several datasets, with all detailed results summarized in Supplementary Table 5.
Validation was performed by reconstructing the MRS in a dataset with available exposure measurements and assessing concordance between MRS values and the corresponding recorded exposure. By leveraging TCGA data, we observed a higher alcohol MRS in liver cancer patients with alcoholic liver disease (ALD), and a separation of current and former smokers from never smokers with the smoking-Maas across several cancer types (Supplementary Table 4 and Extended Data Fig. 1). We further validated the use of MRSs as proxies for cannabis dependence, higher PM2.5 median exposure, and smokers (smoking-Maas) using publicly available datasets, including noncancer participants (Extended Data Fig. 1 and Supplementary Table 5).
Beyond the exposome traits included in our study, we examined additional traits using EWAS and publicly available datasets with DNA methylation and measured exposome data, including allergic asthma, major depressive disorder (MDD), polygenic risk scores (PRS) for depression, as well as prenatal exposures, including birthweight, gestational age, maternal education, maternal BMI and arsenic exposure. The constructed MRSs for these traits showed concordance with directly measured exposures, further strengthening the validity of MRSs as a proxy for exposure (Supplementary Table 5 and Extended Data Figs. 2–3).
Exposome-related methylation risk scores in EOCRC versus LOCRC
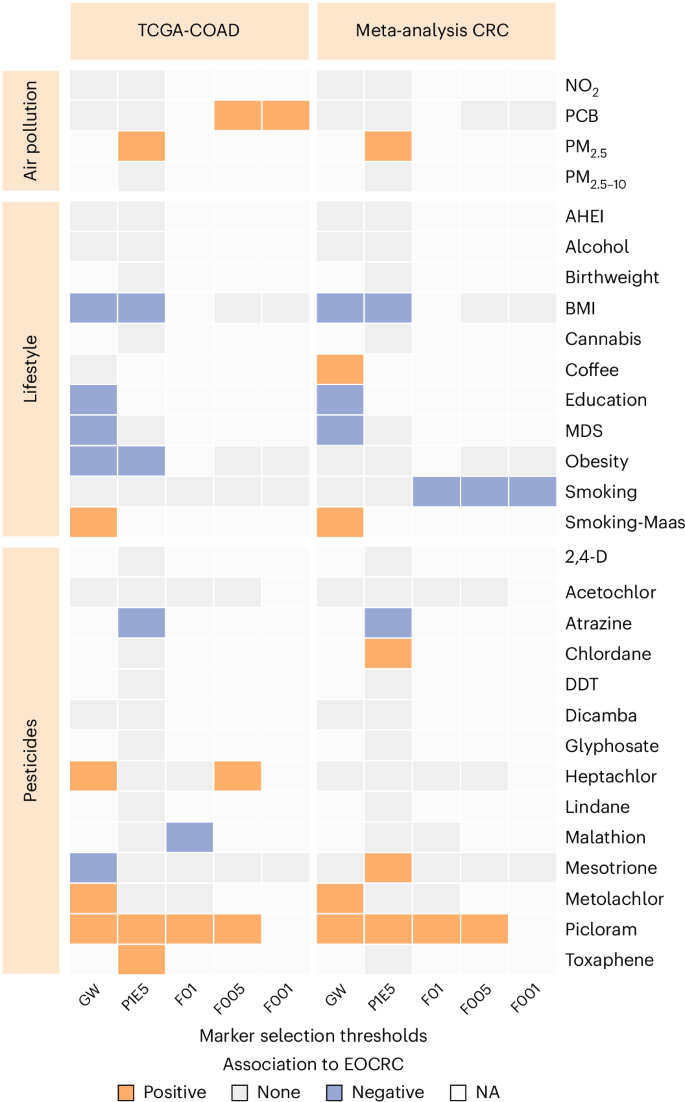
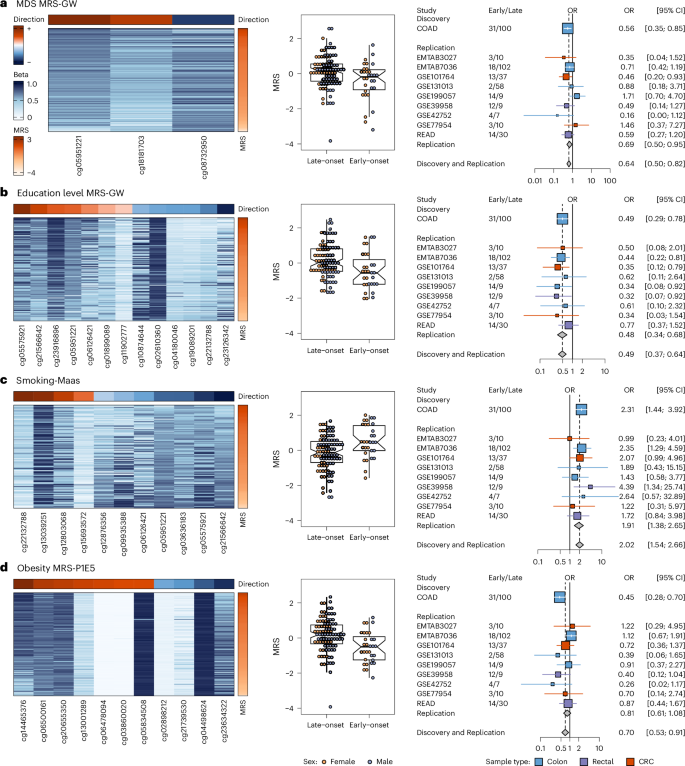
To elucidate the exposome’s impact on early-onset colon and rectal cancer cases, we compared the 63 MRSs between early-onset and late-onset (reference group) patients using multivariate logistic regression models. Due to sex disparities in CRC incidence21, we adjusted the regressions for sex when feasible. Figure 1 summarizes the results for all 29 exposome traits, analyzed across the five selection thresholds used in the original EWAS studies. In the discovery dataset, positive associations were observed for MRSs related to PCB, PM2.5, smoking-Maas, heptachlor, metolachlor, picloram and toxaphene. In contrast, negative associations were found for MRSs corresponding to BMI, education level, MDS, obesity, atrazine, malathion and mesotrione (Fig. 1 and Supplementary Table 6). We highlight the results for four lifestyle factors previously linked to colon and rectal cancers, including the MDS22 (Fig. 2a) and education level23 (Fig. 2b), which are considered protective factors, as well as smoking habits24 (Fig. 2c) and obesity status25 (Fig. 2d), which are recognized as risk factors. To examine the directionality of our findings, the heatmaps in the left panels of Fig. 2 show the methylation level distributions across CpGs featured in each of the four MRSs, along with their direction in the original EWAS and after sorting by the derived MRSs. The heatmaps show that increased MRS correlates with higher beta-values in CpGs with positive associations in the EWAS and with lower beta-values in CpGs with negative associations (see Extended Data Fig. 4 for a more detailed explanation). These results suggest that an elevated MRS reflects greater exposure levels in the original EWAS. Specifically, for patients with early-onset colon cancer, this suggests a lower MDS (P = 7.9 × 10−3; adjusted P (Padj.) = 3.3 × 10−2) (Fig. 2a), lower education levels (P = 4.0 × 10−3; Padj. = 2.2 × 10−2) (Fig. 2b), increased smoking (smoking-Maas) exposure (P = 1.1 × 10−3; Padj. = 8.9 × 10−3) (Fig. 2c), and lower obesity rates (P = 8.4 × 10−4; Padj. = 7.5 × 10−3) (Fig. 2d) in comparison to those with late-onset, as illustrated in the middle panels of Fig. 2. The association of lower obesity rates with early-onset cases was verified utilizing physical metrics from TCGA-COAD. Colon cancer patients with a BMI over 30 kg m−2, as measured in the clinic, were categorized as obese, resulting in 4 out of 24 early-onset and 18 out of 72 later-onset patients being marked as obese. This provides a relative risk (RR) of 0.67 (95% confidence interval (CI): 0.26–1.76) for obesity in early-onset colon cancer patients within the TCGA-COAD cohort (Extended Data Fig. 5), supporting the MRS results for obesity.
Summary overview of the association between the 63 exposome-related MRSs across 29 exposome traits (y axis) and five selection thresholds (x axis, as applied in the original EWAS), comparing early-onset (age <50 years) to late-onset (age ≥70 years, reference) cancer using sex-adjusted logistic regression. The discovery phase (left panel) was conducted in TCGA-COAD (31 early-onset versus 100 late-onset colon cancer). The replication phase (right panel) shows meta-analysis results from nine cohorts (E-MTAB-3027, E-MTAB-7036, GSE199057, GSE131013, GSE42752, TCGA-READ, GSE39958, GSE101764 and GSE77954), including 83 EOCRC and 272 LOCRC patients. The selection thresholds (x axis) indicate the significance threshold in the original EWAS used for CpG selection, namely P < 1.2 × 10−7 (GW), P < 1.0 × 10−5 (P1E5) and FDRs of <0.1 (F01), <0.05 (F005) and <00.1 (F001). The presence of CpGs across the five selection thresholds varies, with unavailable data indicated in white (NA), and nonsignificant findings in light gray.
a–d, Lifestyle-related MRS differed between EOCRC and LOCRC, including non-Mediterranean dietary patterns MRS-GW (marker threshold P < 1.2 × 10−7) (a), lower education levels MRS-GW (b), higher smoking exposure (smoking-Maas) (c) and lower obesity rates MRS-P1E5 (marker threshold P < 1.0 × 10−5) (d) in patients with EOCRC. The heatmaps (left) display epigenetic age-adjusted DNA methylation beta-values in TCGA-COAD, ordered by MRS; the top color bar indicates CpG effect direction in the original EWAS (red: positive; blue: negative). The boxplots (middle) show MRS distribution in patients with EOCRC (N = 31) and LOCRC (N = 100) in TCGA-COAD, stratified by sex (orange: female; purple: male). The forest plots (right) present sex-adjusted logistic regression results for each replication dataset, their meta-analysis (Replication), and combined with TCGA-COAD (Discovery and Replication), for colon (blue), rectal (purple) and CRC (red). Squares: ORs; horizontal lines: 95% CIs. Boxplots show median (center line), interquartile range (IQR) (box) and whiskers (1.5× IQR) and outliers.
Extending our investigation, we conducted a meta-analysis across the nine replication datasets (Figs. 1 and 2 and Supplementary Table 7). This analysis corroborated the initial findings, notably the associations of a lower MDS (P = 1.5×10−2; Padj. = 4.6×10−2), lower educational levels (P = 2.11 × 10−5; Padj. = 3.59 × 10−4) and higher smoking exposure (P = 1.02 × 10−5; Padj. = 8.6 × 10−4) in EOCRC (Fig. 2, right panel). To ensure platform consistency, the meta-analysis was repeated using eight datasets generated with the 450K array, excluding the EPIC-based cohort GSE199057. This strengthened the association for lower MDS (P = 3.06 × 10−3; Padj. = 1.3 × 10−2), whereas educational level and smoking-Maas remained significant (Supplementary Table 8). Finally, despite the considerably reduced sample size, an exploratory tissue-specific meta-analysis of rectal-only (TCGA-READ and GSE39958) or colon-only cancer datasets (GSE131013, GSE42752, E-MTAB-7036 and GSE199057) showed a consistent trend in colon for education and smoking (Extended Data Fig. 6 and Supplementary Table 9).
Picloram-related MRSs
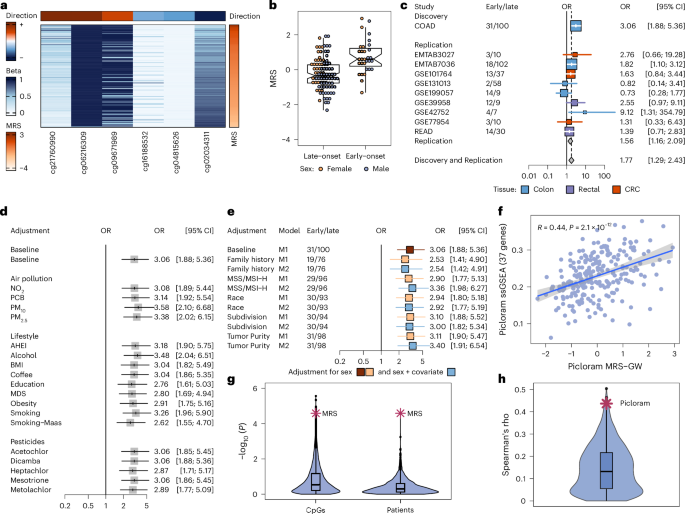
Our results highlight a new association between the MRSs for the herbicide picloram and EOCRC, in comparison to LOCRC cases, in both the discovery and meta-analysis (Fig. 1). We observed that a higher exposure level, as indicated by the original EWAS direction, is associated with an elevated MRS (Fig. 3a). This association highlights increased exposure to picloram among patients with early-onset colon cancer (P = 2.59 × 10−5; Padj. = 4.41 × 10−4) (Fig. 3b)—a finding consistently supported by our meta-analysis using all replication cohorts (P = 3.07 × 10−3; Padj. = 1.51 × 10−2; odds ratio (OR): 1.56 [95% CI: 1.16–2.09]) (Fig. 3c) as well as in the meta-analysis using only 450K array samples (P = 7.17 × 10−4; Padj. = 4.06 × 10−3; OR: 1.71 [95% CI: 1.25–2.33]).
The figure shows the distribution and validation of the picloram MRS employing the genome-wide CpG selection threshold (MRS-GW). a, Heatmap of epigenetic age-adjusted DNA methylation beta-values in TCGA-COAD, ordered by MRS; the top color bar indicates CpG effect direction in the original EWAS. b, Boxplot showing MRS-GW distribution in early- (N = 31) and late- (N = 100) onset TCGA-COAD patients, stratified by sex (orange, female; purple, male). c, Forest plot showing sex-adjusted logistic regression results in replication cohorts, their meta-analysis (Replication), and combined discovery and replication, including colon (blue), rectal (purple) and colorectal (red) cancer. d,e, The associations were further adjusted for other MRSs (31 EOCRC versus 100 LOCRC) (d) and tumor purity, subdivision (left/right), first-degree family with cancer diagnosis (yes/no), race (White/non-White) and MSI status (e). The baseline model (red) includes sex adjustment only; subset analyses show sex-adjusted (M1, orange) and fully adjusted models (M2, blue square). f, Spearman correlation between the 37-gene picloram-specific ssGSEA and the picloram MRS in TCGA-COAD (P = 2.127845 × 10−12); solid line: linear fit with the shaded area indicating the 95% CI. g,h, The violin plot shows the permutation results, confirming the robustness of CpG selection, patient categorization (g) and ssGSEA association (h). In the forest plots, squares represent ORs, and lines 95% CIs. Boxplots show median (center line), IQR (box), whiskers (1.5× IQR) and outliers.
The model presented was initially adjusted only for sex to facilitate external validation in independent datasets lacking measurements for additional variables. To evaluate the robustness of the association between picloram MRS-GW and age at onset, we applied additional adjustments in TCGA-COAD patients. First, we accounted for the influence of other MRSs included in the GW marker selection threshold (Fig. 3d). We observed only minor variations across most adjustments, and none resulted in the loss of the significant association between picloram MRS-GW and age at onset (Fig. 3d). Second, we explored the distribution of the picloram MRS-GW between male and female participants (P = 0.0024) and between microsatellite stable (MSS) and instable (MSI) tumors (P = 0.01), tumor purity high (≥0.7) versus low (<0.7) (P = 5.3 × 10−15), and across the consensus molecular subtypes (Extended Data Fig. 7). The observed significant differences highlight the need for further adjustment when testing the association between picloram MRS-GW and age at onset. As not all variables were available for every patient, Fig. 3e presents Model 1 (adjusted only for sex) alongside Model 2, which incorporates each additional adjustment. This approach ensures consistency by analyzing the same subset of patients to account for sample size differences. Notably, no substantial changes in the association between picloram MRS-GW and age at onset were observed after adjustments.
Picloram-induced gene expression alterations are associated with picloram MRS-GW
A high picloram MRS-GW score showed robust associations with EOCRC patients; however, validating whether the picloram MRS reflects actual picloram exposure is not feasible, as data on picloram-related DNA methylation effects are, to our knowledge, unavailable. Alternatively, we validated the pesticide MRSs using changes in gene expression induced by pesticides. Our rationale was that if the gene expression changes induced by pesticide exposure are reflected in the respective MRS, this would support that the MRS represents the molecular effects of pesticide exposure. To this end, we analyzed RNA-seq data (GSE262419) from induced pluripotent stem cell (iPSC)-derived cardiomyocytes exposed to a wide range of chemicals (N = 67) including several pesticides overlapping with those in our study. To identify gene expression alterations for each pesticide, we performed differential gene expression analysis (DGEA) using pesticide exposure as a continuous variable (0.2 µM, 1 µM and 10 µM). Differentially expressed genes (P < 0.005) were then used to generate single-sample gene set enrichment analysis (ssGSEA) scores in TCGA-COAD. The Spearman correlation between each pesticide’s ssGSEA score and its corresponding MRS was tested, identifying notable correlations for dicamba (six genes: R = −0.15; P = 0.025), heptachlor (148 genes: R = −0.15; P = 0.017) and picloram (37 genes: R = 0.44; P = 2 × 10−12) (Fig. 3f), whereas nonsignificant trends were obtained for acetochlor (16 genes: R = −0.015; P = 0.81), atrazine (90 genes; R = −0.07; P = 0.28), lindane (38 genes: R = 0.002, P = 0.98) and metolachlor (51 genes; R = 0.049, P = 0.46). The observed correlations between picloram and dicamba-specific MRSs and ssGSEA scores suggest that these MRSs capture molecular footprints of exposure, supporting their potential as proxies for true pesticide exposure.
Robustness testing for picloram CpGs, patient categorization and the ssGSEA score
To assess robustness, we performed permutation tests to ensure that the observed associations stem from biological relationships rather than artifacts of CpG selection, patient classification or gene selection. CpGs in the picloram MRS-GW ranked 13th in significance among 10,000 permutations (Fig. 3g). Furthermore, patient classification permutation identified age-based classification as the second most significant, based on picloram MRS-GW, among 1,000 permutations (Fig. 3g). CpG site permutations for MDS, education levels, smoking-Maas and obesity are described in Extended Data Fig. 8a and onset categorization permutations in Extended Data Fig. 8b. Finally, permutation testing ranked the picloram-specific ssGSEA score 6th among 1,000 permutations (Fig. 3h), whereas dicamba and heptachlor ranked 71st and 300th, respectively. These results confirm the robustness of CpGs in picloram MRS-GW, patient categorization, and the genes included in the picloram and dicamba ssGSEA scores.
Molecular differences between high and low picloram MRS-GW
Given the robust association between the picloram MRS-GW and younger age at diagnosis, we investigated its relationship with tumor mutations and gene expression. Comparing tumors in the lowest and highest picloram MRS-GW terciles, we identified 75 differentially mutated genes (P < 0.05) in the full dataset, including 42 mutated only in the high exposure (Extended Data Fig. 9). Due to known differences between MSS and MSI, we performed a stratified analysis (Supplementary Table 10). In MSS tumors (Fig. 4a), eight genes were mutated differentially, with one gene restricted to high picloram exposure and five genes restricted to low exposure. Finally, in MSI tumors, 25 genes were mutated differentially (Fig. 4c), including 11 genes unique to low exposure and 10 genes unique to high exposure. The mutation distribution of the identified genes is presented in Fig. 4a–d.
a,b, Tumors were divided into terciles of the picloram MRS-GW, and tumors in the first and third terciles were compared using Fisher’s exact test to identify differentially mutated (P < 0.05) genes. Differentially mutated genes in MSS tumors are shown with the percentage of tumors with a gene mutation in the first or third tercile (a) and their distribution across all MSS tumors, sorted on the picloram MRS (b). c,d, Similarly, the differentially mutated genes in MSI tumors are shown with the percentage of tumors having the gene mutated in the first or third tercile (c) and their distribution across all MSI tumors (d). e, Comparison of gene expression between tumors in the first and third tercile of the picloram MRS-GW using negative binomial generalized linear models. The analysis was adjusted for age, sex, MSI status, tumor purity and tumor location (left/right) and significance thresholds were set at an absolute log2FC > 1.2 and a Padj. < 0.05, resulting in 25 differentially expressed genes with the top 6 genes highlighted. f, NESs for the significantly down- and upregulated pathways in low picloram exposure, using the differential expression results.
DGEA adjusted for age, sex, MSI status, tumor purity and tumor location (left/right) identified 25 differentially expressed genes (Padj. < 0.05, |log2fold-change (FC) | > 1.2), including 16 downregulated genes in low picloram exposure (Fig. 4c and Supplementary Table 11). MSS-stratified analysis identified four differentially expressed genes, whereas MSI-stratified analysis identified 244 genes (Supplementary Table 11). The subsequent GSEA using all samples revealed nine enriched pathways (Padj. < 0.05, |normalized enrichment score (NES)| > 1), including upregulation of the Wnt/β-catenin signaling in low picloram exposure, consistent with APC mutations and downregulation of immune response-related pathways, such as complement and IL6-JAK-STAT3 signaling.
Young tumors associated with picloram exposure
Current CRC classification into early- or late-onset relies on age at diagnosis; however, variability in the interval between tumor initiation and diagnosis makes age at diagnosis an unreliable indicator of the tumor’s actual age26. We therefore evaluated single-base substitution signature 1 (SBS1), a mitotic division-related mutational signature, as a proxy for tumor age. SBS1 reflects the cumulative number of mitotic divisions and captures the replicative history of cells across premalignant and malignant stages27,28. We used the SBS1 score from the Catalogue of Somatic Mutations in Cancer, deconvolved by the Pan-Cancer Analysis of Whole Genomes consortium using established methods with high accuracy (≥0.827)28, and no samples were excluded based on accuracy (Extended Data Fig. 10a). TCGA-COAD patients with DNA methylation and mutational signatures data were included, excluding MSI tumors due to distinct mutational mechanisms (Extended Data Fig. 10b)29,30. SBS1 distribution across age groups early-, intermediate- (IOCRC: aged 50–69 years) and late- (N = 173) onset is detailed in Extended Data Fig. 10c.
Picloram MRS-GW was associated significantly with early-onset versus late-onset cases (N = 25 versus 72; OR: 2.99 [95% CI: 1.70–5.85]; P = 4.27 × 10−4). Next, we employed an SBS1 score threshold, identifying 72 young (SBS1 < 60) and 101 old tumors (SBS1 ≥ 60). The new patient categorization underscored a significant association with picloram MRS-GW (OR: 1.84 [95% CI: 1.31–2.66]; P = 6.57 × 10−4) (Extended Data Fig. 10d). The chronological age distribution for these SBS1-defined groups is shown in Extended Data Fig. 10e. This association remained significant using a non-age-adjusted MRS-GW when adjusting for sex (P = 4.15 × 10−4) and further for age at diagnosis (P = 5.98 × 10−3), confirming independence from chronological age. The SBS1 score was associated strongly with tumor mutation burden (TMB; R = 0.66, P < 2.2 × 10−16). Consequently, adjusting the original model (P = 6.57 × 10−4) for TMB attenuated the association (P = 3.23 × 10−2). To further investigate whether TMB could serve as a marker for tumor age, we performed linear regression models adjusted for sex, showing a stronger association between the picloram MRS-GW and SBS1 score (P = 2.11 × 10−5) than TMB (P = 0.0148), suggesting that although TMB and SBS1 are highly correlated, SBS1 captures the age-specific signal more effectively.
County-level pesticide uses and EOCRC incidence across the USA
Using MRSs as proxies, we identified associations between age at onset and several pesticides. We examined pesticide use intensity and EOCRC incidence rates (IR) across overlapping counties in California, Connecticut, Georgia, Iowa, New Mexico, Utah and Washington in the USA. Pesticide use estimates (1992–2012) were extracted from the Pesticide National Synthesis Project, and the age-adjusted EOCRC IR from the Surveillance, Epidemiology and End Results (SEER) for eight registries (SEER8). Among 225 pesticides with sufficient data (≥10 years and ≥50 total observations), 62 were associated with EOCRC IR in linear mixed models adjusted for years of data collection and county-level random effects (Supplementary Table 12). After additional adjustment for seven county-level socioeconomic status (SES) indicators31, including median household income, median house value, median rent, percent below 150% of the poverty line, education index32, percent working class and percent unemployed, 27 pesticides remained significant (Fig. 5a and Supplementary Table 13).
County-level pesticide use intensity estimates (1992–2012) were matched with EOCRC incidence rates. a, Heatmap showing associations between 62 pesticides (y axis) and EOCRC IRs, adjusted for seven SES indicators (x axis). b, The 27 pesticides (y axis) that remained significant were further adjusted for each of the other 26 pesticides (x axis). Pesticides that lost significance in the baseline model without pesticide adjustment are shown in dark pink, whereas those that were significant in the basic model but lost significance after pesticide adjustment are displayed in light pink. Cases with insufficient datapoints for testing are shown in gray. c, The five most robust pesticides were analyzed jointly, adjusting for the remaining pesticides and SES indicators. The left y axis represents the top five pesticides, including atrazine, esfenvalerate, glyphosate, nicosulfuron and picloram, whereas the right y axis reflects for each of them the four pesticides they are adjusted for. The x axis shows the adjustments for only the pesticide or the pesticide and each of the SES indicators. NS, nonsignificant.
As several pesticides were used within counties, we tested robustness by adjusting each of the 27 pesticides for the remaining 26 (Fig. 5b and Supplementary Table 13). Since merging the pesticide data can reduce the number of overlapping datapoints substantially, we first tested baseline models without pesticide adjustment to distinguish true attenuation from reduced data overlap. Reduced overlap led to frequent loss of significance, particularly for fenbuconazole (0.84), propargite (0.80) and flumioxazin (0.65) (Fig. 5b). Among the remaining 24 pesticides, glyphosate and picloram remained significant in 21 of 23 pairwise models, esfenvalerate and nicosulfuron in 15, and atrazine in 12, with others remaining significant in ten or fewer models.
Integrated pesticide- and SES-adjusted analyses of the five most robust pesticides showed limited consistency for atrazine and esfenvalerate, each significant in only one pesticide-adjusted model and moderate robustness for nicosulfuron, which remained significant in two models. Glyphosate showed stronger robustness, maintaining significance in three models, including two with all SES adjustments. Picloram demonstrated the most consistent association, remaining significant in all four pesticide-adjusted models and across most SES-adjusted models (Fig. 5c and Supplementary Table 14). Together with the MRS findings, these results highlight picloram as a potentially important environmental factor contributing to EOCRC risk.
Discussion
In this study, through the use of epigenetic markers, we explored the exposome traits that could be contributing specifically to EOCRC disease. A principal challenge in exposome research is the limited availability of direct exposure measurements, making it difficult to fully reconstruct a person’s exposome profile. This research addresses the exposome impact, including environmental and lifestyle factors, on early-onset colon and rectal cancer cases through epigenetic fingerprints.
Despite the limitations in exposome data availability and the possibility of confounding bias, we validated the use of MRSs as a proxy for exposome exposure using datasets containing direct exposome measurements. In the context of alcohol consumption, we used the presence of ALD as a surrogate marker and showed that patients with ALD had higher alcohol MRS-GW scores compared to those without ALD. Regarding smoking exposure, our findings indicate that once the smoking-related methylation signature becomes embedded in tumor tissue it remains persistent and irreversible, even after prolonged (>15 years) smoking cessation. This highlights the lasting epigenetic imprint of smoking exposure on tumor DNA.
Exposures across the lifespan may change, so the analyses incorporating both categorical and quantitative measurements are challenging and may raise concerns about potential spurious associations. Our results consistently link EOCRC to exposure to the herbicide picloram based on epigenetic fingerprints. Picloram’s mechanism of action as an herbicide relies on its capacity to mimic the plant growth hormones auxins and to inhibit the enzymes that break down auxins, which leads to more persistent effects than the natural hormone. Therefore, picloram disrupts normal growth, causing abnormal stimulation and maturation of tissues, which triggers growth discontinuation, root deterioration and eventually plant death. Picloram was first registered as an herbicide in the USA in 1964, and the herbicide and its derivatives have generally shown to be of moderate-to-low acute toxicity in laboratory animals33. However, dietary exposure to residues of picloram is plausible as it has been found in grain and meat byproducts, and the effects of long-term use on human health have not been described so far. If the use of picloram in crops started in the mid and late twentieth century, then current patients with LOCRC were not exposed during their childhood, whereas cases of EOCRC were and have been for a longer part of their lives, which could explain our results. The association between pesticide exposure and EOCRC using population-based data further validates the association with picloram.
Distinct molecular aberrations are observed when comparing tumors in the first tertile of the picloram MRS to those in the third tertile. Specifically, we identified an upregulation of the Wnt/β-catenin signaling pathway and a higher prevalence of APC mutations (90%) in low picloram tumors compared to 74% in high picloram tumors. These findings suggest that picloram, along with other exposome factors, may influence tumor development by promoting alternative oncogenic pathways, diverging from the classical Fearon and Vogelstein model34. Nevertheless, given the scarcity of studies investigating picloram’s impact, particularly on DNA methylation and health outcomes in humans, we emphasize the urgent need for longitudinal and experimental research. We believe that determining the reversibility of picloram-induced epigenetic changes in normal tissue and tumors, establishing a causal link between picloram exposure and colorectal carcinogenesis, and understanding its dose-dependent risk are critical next steps in disentangling the EOCRC epidemic.
Besides picloram, we also show evidence for associations between EOCRC and exposure to glyphosate, esfenvalerate, nicosulfuron and atrazine. In the USA, atrazine is one of the most widely used herbicides—as a selective herbicide to prevent weeds from germinating in corn production—whereas glyphosate is applied extensively in corn, soybeans and other crops as a broad-spectrum herbicide used to eliminate weeds that have already emerged, creating a complementary weed control strategy. Glyphosate is already categorized as ‘probably carcinogenic to humans’ by the International Agency for Research on Cancer (IARC), suggesting the validity of the obtained results35. Although atrazine has been banned in the European Union since 2004, the United States Environmental Protection Agency still approves its continued use36. New studies have identified several mechanisms for how atrazine could cause carcinogenesis in humans, for example, by damaging DNA integrity, the stability of the cell genome, DNA double-strand breaks and the activation of DNA damage checkpoints37. This resulted in the Monographs program for 2025–202938 to warrant a high-priority re-evaluation of the IACR classification as a result of the newly identified human cancer and mechanistic evidence. Furthermore, a recent update study in the Agricultural Health Study Cohort further suggested the link between atrazine use and several cancer types, including in patients <50 years of age39.
Lower education levels were found for patients with EOCRC. Educational attainment is an important determinant of health and lifespan40 and it is used frequently as a proxy for SES in health studies, revealing strong associations with the prevalence of noncommunicable diseases, including cancer41. Furthermore, findings from the International Public Opinion Survey on Cancer 202042 highlight that people without a university degree are less aware of modifiable cancer risk factors compared to those with higher educational attainment. These findings suggest that education level not only influences health outcomes but also plays a critical role in shaping awareness and the adoption of healthier lifestyle behaviors.
The patient’s age at diagnosis is currently used as the tumor’s age, possibly introducing bias in identifying early-onset patients. To address this, we assessed whether tumor age can be established by identifying molecular characteristics of aged tumors, such as mutational signatures27,28. Characteristic mutational signatures in cancer genomes arise from various mutational processes, including defects in DNA maintenance mechanisms and both external and internal exposures28. Mutations attributed to SBS1 are thought to occur during DNA replication in mitosis, suggesting that the rate of SBS1 mutations may serve as an indicator of the number of mitotic divisions a cell has undergone27,28. Furthermore, the number of SBS1-attributable mutations within a tumor is correlated with patient age at cancer diagnosis27,28. Here we demonstrated that using an SBS1 score threshold to distinguish between younger and older tumors is associated with picloram MRS. This finding suggests that the observed differences may be attributable to tumor biological age rather than the patient’s chronological age.
We acknowledge the limitations of the current study. First, the number of EOCRC cases in the datasets is relatively small, specifically in the analysis of colon and rectal cancers separately. We emphasize the need for larger studies to capture the diversity of worldwide patients with EOCRC in terms of ethnicity, cultural and socioeconomic differences. Another limitation of this study is the restricted availability of pesticide exposure data, which began only in 1992, as well as the lack of data on tumor onset age, with only age at diagnosis available. Because of the substantial time lag between onset and diagnosis, these constraints prevent us from assessing accurately the latency period between picloram exposure and EOCRC development. This emphasizes the need for future in vivo studies to examine the causality and latency period of picloram in EOCRC.
The study also presents unprecedented strengths. First, the use of MRS as a proxy for exposome factors is a new approach that enables the exploration of traits that would otherwise be unfeasible due to limited data availability. Similarly, the lack of longitudinal follow-up measurements or reliance on self-reported data can be overcome, as MRSs integrate the cumulative impact of exposure over time. Nevertheless, prospective birth cohorts with long-term follow-up, quantitative exposure measurement, and biomarker and omics analyses throughout life can elucidate the etiology of EOCRC. In addition, given that early-life exposures may be key to CRC development, prospective birth cohorts are needed. Second, several independent cohorts were used, and permutation tests were implemented; the picloram MRS was associated consistently with EOCRC compared with LOCRC. Validation of the association between picloram use intensity and EOCRC in population-level data also further strengthens the evidence for picloram as a new risk factor. Third, the exposome, namely the totality of exposures including, among others, diet, lifestyle and environment, during early life and young adulthood, has changed considerably in the last decades. The categorization of the population into two cohorts with extreme ages (<50 versus ≥70) and the exclusion of cases with intermediate age favor the identification of substantial generational changes in the exposome of the two cohorts, whereas those in between may show a gradual change in their exposure. This investigation has substantial relevance and broad potential impact, as it identifies research priorities for primary interventions through behavior modification and secondary prevention among people exposed to EOCRC risk factors, for current and future generations. Moreover, it might guide the development of health policies for environmental exposures and regulatory guidelines for agricultural products. Finally, the spotlight on the tumor age rather than on the patient is a new perspective for CRC research and epidemiology for both EOCRC and those diagnosed over the age of 50 years. This could be one of the potential explanations for why no differences in tumor biology between EOCRC and LOCRC have been found to date.
In conclusion, our findings not only provide exposome traits based on epigenetic fingerprints that could be contributing to the development of CRC, specifically in EOCRC, but also pave the way with a compelling rationale for addressing lifestyle and environmental exposures to mitigate EOCRC risk, highlighting the importance of both personal and policy-level interventions.
Methods
Study population
The discovery phase of our study utilized TCGA-COAD samples. To ensure the integrity of our analysis, patients with Lynch syndrome (TCGA-A6-6781, TCGA-CM-6674 and TCGA-D5-6927) were excluded. Our replication effort encompassed nine datasets, focusing on rectal (TCGA-READ and GSE39958)15, colon (GSE131013, GSE42752, E-MTAB-7036 and GSE199057)11,12,13,14, and colorectal (GSE101764, GSE77954 and E-MTAB-3027)16,17,18 cancer studies. For the TCGA samples, we implemented exclusion criteria, removing cases annotated with ‘Item in special subset,’ ‘History of unacceptable prior treatment related to a prior/other malignancy,’ ‘Case submitted is found to be a recurrence after submission,’ ‘Neoadjuvant therapy,’ ‘Synchronous malignancy,’ and ‘Pathology outside specification.’ This led to the exclusion of ten patients from the COAD dataset and eight from the READ dataset. Furthermore, samples preserved in formalin-fixed paraffin-embedded form were excluded to maintain consistency in sample quality. Duplicate samples were identified and removed based on their plate number, further refining our cohorts for analysis. Participants were categorized based on age at diagnosis, including early-onset (aged <50 years), intermediate-onset (aged between 50 and 69 years), and late-onset (aged ≥70 years), following the literature for late-onset cut-off43.
DNA methylation data processing
DNA methylation data was in all datasets obtained using the Illumina Infinium HumanMethylation450 BeadChip, except in GSE199057, in which the Illumina Infinium Human Methylation EPIC BeadChip array was used. TCGA methylation data was acquired from the Genomic Data Commons (GDC) Data Portal through the TCGABiolinks package v.2.25.0 (ref. 44). For the additional datasets, idat files or signal intensity files were downloaded from the Gene Expression Omnibus (GEO) database using the GEOquery package v.2.60.0 (ref. 45). We processed the tumor sample methylation data with the sesame package v.1.19.7 and v.1.24.0 (ref. 46) in R v.4.3.1. CpG sites showing more than 50% missing data were removed from each dataset. The remaining missing values underwent imputation using the impute package v.1.66.0 (ref. 47), followed by mean imputation for CpGs with imputed values of 0. DNA methylation beta-values were then converted to M-values using the lumi package v.2.44.0 (ref. 48). Next, we estimated the epigenetic age using the Horvath clock, following the R software tutorial20, and adjusted the DNA methylation m-values for the estimated age to account for the influence of age on methylation. Finally, we converted the m-values back to beta-values for further analysis. The final dataset refinement involved excluding CpGs that were not present across all datasets, those associated with cross-reactive probes and CpGs identified as single-nucleotide polymorphisms49.
Exposome-related traits marker selection
We conducted a detailed literature review to compile a list of exposome traits potentially influencing the risk of EOCRC, ensuring the inclusion of traits with available comprehensive EWAS. This process led to the identification of 29 key exposome traits19,50,51,52,53,54,55,56,57,58,59,60,61,62. The traits analyzed encompassed 11 lifestyle factors: the AHEI50, alcohol consumption51, birth weight52, BMI (continuous variable in kg m−2)53, cannabis use62, coffee consumption54, education level55, MDS50, obesity (defined as ≥30 kg m−2)56, smoking habits57 and smoking-Maas19. Furthermore, we examined four air pollution particles: NO2 (ref. 58), PCBs60 and PM2.5 and PM2.5–10 (ref. 59). In addition, we included 14 pesticides encompassing 2,4-D, atrazine, acetochlor, chlordane, dicamba, malathion, DDT, heptachlor, lindane, glyphosate, mesotrione, metolachlor, picloram and toxaphene61. CpG sites were chosen using stringent significance criteria: P < 1.2 × 10−7, P < 1.0 × 10−5 and FDR of <0.01, <0.05 and <0.1. We employed a tiered strategy for marker selection to balance sensitivity and specificity, facilitating a comprehensive assessment of the relationship between exposome exposures, DNA methylation levels and EOCRC. This approach includes a broad range of CpG sites explored using less stringent P value thresholds (for example, P < 1.0 × 10−5 and FDR < 0.1), increasing sensitivity to include CpG sites with potential biological relationships. This was instrumental in selecting both highly confident associations and those of potential biological relevance that may not meet the strictest statistical thresholds. In addition, we included CpG sites using more stringent criteria (for example, P < 1.2 × 10−7, FDR < 0.05 and FDR < 0.01), enhancing specificity and reducing the risk of including false positive CpG sites. This approach allowed for a nuanced exploration of the data, ensuring that both robust and biologically meaningful associated CpG sites were included in our analysis.
Exposome-related methylation risk scores in patients with EOCRC versus LOCRC
To assess the impact of these exposome traits on EOCRC risk, we computed individual MRS by applying effect sizes from relevant EWAS to methylation levels at each CpG site and summing these across all pertinent CpGs.
The performance of these MRSs as proxy measures was evaluated by reconstructing the scores in a dataset63,64,65,66,67,68,69,70,71,72 with directly measured exposure data and assessing the level of agreement between the derived MRS values and the corresponding observed exposure levels. In addition to the exposome traits included in our study, the analysis was extended to a broader set of exposures to further validate the applicability of MRSs as a proxy, including allergic asthma73, MDD74, PRS for depression75, birthweight76,77, gestational age76, maternal education78, maternal BMI79,80 and arsenic exposure69.
After constructing the scores in the tumor cohorts, the scores were normalized to a mean of 0 and an s.d. of 1, using exclusively data from EOCRC and LOCRC patients. We examined the association between these MRSs across early- versus late-onset cases (reference group) by employing logistic regression analyses adjusted for sex, using the stats R package v.4.1.3. This analysis spanned the discovery dataset TCGA-COAD and extended to nine validation cohorts. Models were excluded from the subsequent meta-analysis if they issued fitting warnings or were unable to estimate the 95% CI. The collective data from the validation cohorts were subjected to a meta-analysis utilizing the metafor R package v.4.4-0 (ref. 81). The P values obtained were FDR82-corrected for the number of traits in each marker selection threshold, including 17 traits in GW, 26 in P1E5, 7 in F01, 8 in F005 and 5 in F001, using the p.adjust function in stats v.4.1.3. The results were then summarized visually in forest plots, offering a clear depiction of the associations between MRS and EOCRC versus late-onset cases.
Pesticide-induced gene expression alterations in iPSC-derived cardiomyocytes
The validity of the pesticide MRSs was assessed using gene expression alterations induced by pesticides in iPSC-derived cardiomyocytes. Raw gene expression counts per plate were extracted from GSE262419 (ref. 83). Each plate included raw read counts for 22,537 probes, which were aggregated at the gene-level to yield 19,703 unique genes. Low-expressed genes were then filtered out per pesticide by excluding those with a mean count of fewer than ten across pesticide-exposed samples and those that did not reach at least ten counts in at least two samples. We conducted DGEA (DESeq2 (v.1.44.0)) per pesticide by comparing the exposure doses as a continuous variable. Pesticide-specific gene sets were constructed using DEGs (P < 0.005). These gene sets were used as input to construct ssGSEA scores using the GSVA R package v.1.42.0 (ref. 84) in TCGA-COAD RNA-seq tumor samples. Genes with expression levels of at least ten in more than 50 samples and a variance greater than 0.01 across samples were retained for downstream analyses in TCGA-COAD. The obtained pesticide-specific ssGSEAs were tested for their Spearman correlation with their respective pesticide-specific MRS.
Robustness testing for picloram MRS-GW, CpG selection, patient categorization and ssGSEA genes
The robustness of the findings for picloram MRS-GW was further examined in patients from TCGA-COAD. Additional TCGA data was acquired from the GDC Data Portal through the TCGABiolinks package v.2.25.0 (ref. 44). First, the association between picloram MRS-GW and EOCRC versus LOCRC was further adjusted for the other MRS in the GW marker selection threshold. Second, we further adjusted the association between picloram MRS-GW and EOCRC versus LOCRC by patient-specific characteristics (race and family history) and tumor-specific factors (MSS, tumor location (left/right), and tumor purity). Third, we evaluated the robustness of the selected CpGs and patient categorization by conducting two permutation analyses, ensuring that the observed associations are not artifacts of the specific CpG selection or patient grouping but reflect underlying biological relationships. The first analysis involved assigning patients randomly to the MRSs. Specifically, we shuffled the patients with EOCRC and LOCRC in the TCGA-COAD dataset 1,000 times, creating alternative configurations of the dataset. For each rearranged dataset, we performed logistic regression analyses, adjusting for sex, to assess the stability of the association between MRSs and early-onset across these permutations. The second analysis targeted the specific assignment of CpGs to their corresponding effect sizes (weights). We temporarily removed the CpGs that were selected initially for each exposome trait to create a pool of potential CpGs. From this pool, we generated 10,000 new sets, each containing an equal number of CpGs as in the original analysis but selected randomly. These randomly chosen CpGs were then used to construct new MRSs for each trait. Subsequently, logistic regression analyses, adjusted for sex, were applied to these permutation-derived MRSs to test the significance of the associations under these randomized conditions. Finally, we explored the robustness of the genes included in the pesticide-specific ssGSEA by generating 1,000 random gene sets using the same number of genes as in each specific gene set and computed 1,000 ssGSEA scores. Next, we tested the Spearman correlation between each score and the picloram MRS-GW.
Molecular characterization between high and low picloram MRS
The possible associated molecular landscape of the picloram MRS-GW was explored by comparing the first and third terciles. First, we tested for mutational differences using maftools R package v.2.8.5 (ref. 85). Second, we conducted a DGEA using the DESeq2 R package v.1.44.0, adjusting for age, sex, MSI status, tumor purity and tumor location (left/right). Stratification analyses were performed separately for MSS and MSI tumors. The obtained results were adjusted for multiple testing using the Benjamini–Hochberg procedure for FDR82. DEGs were defined by an absolute log2 FC > 1.2 and an FDR-adjusted P value < 0.05. Finally, we performed GSEA using the fgsea R package v.1.20.0 (ref. 86). Genes were ranked based on the Wald statistic from DESeq2 output, and the Hallmark gene sets (n = 50) were obtained from the HALLMARK Molecular Signatures Database87. Significantly enriched pathways were defined as those with FDR-adjusted P values < 0.05 and absolute NES > 1.
Young versus older tumors using single-base substitution signature 1
Patients from the TCGA-COAD dataset were selected when presented with comprehensive profiles including DNA methylation data, mutational signatures (https://dcc.icgc.org/releases/PCAWG) and confirmed microsatellite stability. To ensure data integrity, we excluded extreme outlier patients, identified by SBS1 scores exceeding the mean SBS1 score by >3 s.d. The patients were categorized into early- (age <50 years), intermediate- (age 50–69 years), and late- (age ≥70 years) onset cases. Initially, we assessed the differential impact of picloram MRS-GW within this subset between the early- and late-onset groups. Subsequently, we aimed to enhance the specificity of our findings by integrating the concept of tumor age, as defined by the SBS1 score28, into our analysis of early- and late-onset categories. In a new approach, we explored the efficacy of using tumor age—defined by an SBS1 score of 60 as a threshold—as an alternative to chronological age in distinguishing between young and old tumors. We selected an SBS1 score threshold of 60 because it corresponded to the first quartile in late-onset patients and the median in intermediate-onset patients. The third quartile in early-onset patients had an SBS1 score of 67.
To ensure that the results observed were not confounded by age at diagnosis, we constructed the picloram MRS-GW using non-age-adjusted DNA methylation levels. Subsequently, we tested its association with young versus old tumors while incorporating an additional adjustment for age at diagnosis. We then explored whether total TMB could serve as a stronger indicator of tumor age than the SBS1 score. TMB was calculated using nonsilent variants and normalized by dividing by 38, corresponding to the standard exome size using maftools R package v.2.8.5 (ref. 85). To evaluate these relationships, we performed linear regression models, assessing the association between picloram MRS-GW and either (1) the SBS1 score or (2) TMB, with both models adjusted for sex.
Pesticide use and EOCRC in the USA
The results obtained for pesticide MRSs were validated using population data from the USA. We assessed the association between pesticide use and EOCRC incidence across overlapping counties. Using data from the Pesticide National Synthesis Project between 1992 and 2012, we analyzed county-level pesticide use intensities by dividing the total estimated pesticide use by the county’s area in square miles.
Age-adjusted IR for colon and rectum (Site and Morphology; Site recode ICD-O-3/WHO 2008: ‘colon and rectum’ without excluding specific tumors) were extracted from the SEER database using the SEER*Stat software (v.8.4.1)88. This comprehensive dataset allowed us to incorporate Research Plus Data spanning from 1975 to 2020 for eight registries (SEER8). We calculated age-adjusted EOCRC incidence rates (for people aged 25–49 years) at the county level annually, expressed as cases per 100,000 population. Counties with no cases for more than 50% of the years were excluded to mitigate the impact of sparse data on our analysis.
The pesticide data and EOCRC incidence data were overlapped at county level per year. To refine our analysis and reduce the influence of extreme values, we removed the top 5% measurements based on log-transformed pesticide use intensity. We used a linear mixed model to examine the relationship between pesticide use intensity and EOCRC incidence, adjusting for the years measured and including a random effect to account for variations across counties, using the lmerTest R package v.3.1-3 (ref. 89). For the pesticides significantly associated with EOCRC IR, we further adjusted the model for seven county-level socioeconomic factors: median household income, median house value, median rent, percentage below 150% of the poverty line, education index, percentage of the working class and percentage unemployed. The pesticides that remained associated significantly were further adjusted for the remaining pesticides. Due to limited overlapping datapoints, we curated datasets for each pesticide combination to maximize the number of observations. If the combination of data for one pesticide with other pesticides was not significant at ≥65%, the pesticide was removed from further analysis. Pesticides that remained significant in ≥50% of the pairwise adjustments were further adjusted for the other pesticides, combined with adjustments for SES, to examine the potential confounding effects of simultaneous pesticide use and SES.
Statistical analyses were performed in R v.4.1.3 (ref. 90).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All methylation and related clinical data used in this study are publicly available via the GDC Data portal (https://portal.gdc.cancer.gov/) for TCGA datasets, the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) for GSE131013, GSE42752, GSE39958, GSE101764, GSE77954, GSE199057, GSE262419, GSE50660, GSE72680, GSE289816, GSE294810, GSE269103, GSE198904, GSE62924, GSE71678, GSE110828, GSE75248 and GSE224363 or the BioStudies database (https://www.ebi.ac.uk/biostudies/) for E-MTAB-7036 and E-MTAB-3027. Pesticide use data can be extracted from the National Water-Quality Assessment (NAWQA) project (https://water.usgs.gov/nawqa/pnsp/usage/maps/county-level/). Access to the Research Plus Data from the SEER cohort can be requested via https://seer.cancer.gov/data/access.html.
Code availability
The code used for the analysis in this study is publicly available via GitHub at https://github.com/CancerCompBioLab/EOCRCexposome (ref. 91). All the input files needed to replicate our findings and the results obtained during the study are also available from the GitHub repository.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024).
Siegel, R. L. et al. Global patterns and trends in colorectal cancer incidence in young adults. Gut 68, 2179–2185 (2019).
Berrington de Gonzalez, A. et al. Trends in cancer incidence in younger and older adults: an international comparative analysis. Ann. Intern. Med. 178, 1677–1687 (2025).
Akimoto, N. et al. Rising incidence of early-onset colorectal cancer—a call to action. Nat. Rev. Clin. Oncol. 18, 230–243 (2021).
Li, J. et al. Patterns in genomic mutations among patients with early-onset colorectal cancer: an international, multicohort, observational study. Lancet Oncol. 26, 1055–1066 (2025).
Syed, A. R. et al. Old vs new: risk factors predicting early onset colorectal cancer. World J. Gastrointest. Oncol. 11, 1011–1020 (2019).
Connell, L. C., Mota, J. M., Braghiroli, M. I. & Hoff, P. M. The rising incidence of younger patients with colorectal cancer: questions about screening, biology, and treatment. Curr. Treat. Options Oncol. 18, 23 (2017).
Wu, H., Eckhardt, C. M. & Baccarelli, A. A. Molecular mechanisms of environmental exposures and human disease. Nat. Rev. Genet. 24, 332–344 (2023).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Díez-Villanueva, A. et al. DNA methylation events in transcription factors and gene expression changes in colon cancer. Epigenomics 12, 1593–1610 (2020).
Naumov, V. A. et al. Genome-scale analysis of DNA methylation in colorectal cancer using Infinium HumanMethylation450 BeadChips. Epigenetics 8, 921–934 (2013).
Fennell, L. et al. Integrative genome-scale DNA methylation analysis of a large and unselected cohort reveals 5 distinct subtypes of colorectal adenocarcinomas. Cell Mol. Gastroenterol. Hepatol. 8, 269–290 (2019).
Ghosh, J. et al. Epigenome-wide study identifies epigenetic outliers in normal mucosa of patients with colorectal cancer. Cancer Prev. Res. (Phila.) 15, 755–766 (2022).
Ha, Y. J. et al. Epigenetic regulation of KLHL34 predictive of pathologic response to preoperative chemoradiation therapy in rectal cancer patients. Int. J. Radiat. Oncol. Biol. Phys. 91, 650–658 (2015).
Barrow, T. M. et al. Smoking is associated with hypermethylation of the APC 1A promoter in colorectal cancer: the ColoCare Study. J. Pathol. 243, 366–375 (2017).
Qu, X. et al. Integrated genomic analysis of colorectal cancer progression reveals activation of EGFR through demethylation of the EREG promoter. Oncogene 35, 6403–6415 (2016).
Lennard, K. S., Goosen, R. W. & Blackburn, J. M. Bacterially-associated transcriptional remodelling in a distinct genomic subtype of colorectal cancer provides a plausible molecular basis for disease development. PLoS ONE 11, e0166282 (2016).
Maas, S. C. E. et al. Validated inference of smoking habits from blood with a finite DNA methylation marker set. Eur. J. Epidemiol. 34, 1055–1074 (2019).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Ramai, D. et al. Gender and racial disparities in colorectal cancer incidence and mortality: a national cancer registry study. Int. J. Colorectal Dis. 36, 1801–1804 (2021).
Zhong, Y. et al. Association between Mediterranean diet adherence and colorectal cancer: a dose-response meta-analysis. Am. J. Clin. Nutr. 111, 1214–1225 (2020).
Doubeni, C. A. et al. Socioeconomic status and the risk of colorectal cancer: an analysis of more than a half million adults in the National Institutes of Health-AARP Diet and Health Study. Cancer 118, 3636–3644 (2012).
Botteri, E. et al. Smoking and colorectal cancer risk, overall and by molecular subtypes: a meta-analysis. Am. J. Gastroenterol. 115, 1940–1949 (2020).
Mandic, M., Safizadeh, F., Niedermaier, T., Hoffmeister, M. & Brenner, H. Association of overweight, obesity, and recent weight loss with colorectal cancer risk. JAMA Netw Open 6, e239556 (2023).
Umar, A., Dunn, B. K. & Greenwald, P. Future directions in cancer prevention. Nat. Rev. Cancer 12, 835–848 (2012).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Sia, E. A., Kokoska, R. J., Dominska, M., Greenwell, P. & Petes, T. D. Microsatellite instability in yeast: dependence on repeat unit size and DNA mismatch repair genes. Mol. Cell Biol. 17, 2851–2858 (1997).
Boland, C. R. & Goel, A. Microsatellite instability in colorectal cancer. Gastroenterology 138, 2073–2087 (2010).
Yost, K., Perkins, C., Cohen, R., Morris, C. & Wright, W. Socioeconomic status and breast cancer incidence in California for different race/ethnic groups. Cancer Causes Control. 12, 703–711 (2001).
Liu, L., Deapen, D. & Bernstein, L. Socioeconomic status and cancers of the female breast and reproductive organs: a comparison across racial/ethnic populations in Los Angeles County, California (United States). Cancer Causes Control 9, 369–380 (1998).
Reuber, M. D. Carcinogenicity of picloram. J. Toxicol. Environ. Health 7, 207–222 (1981).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Davoren, M. J. & Schiestl, R. H. Glyphosate-based herbicides and cancer risk: a post-IARC decision review of potential mechanisms, policy and avenues of research. Carcinogenesis 39, 1207–1215 (2018).
Sass, J. B. & Colangelo, A. European Union bans atrazine, while the United States negotiates continued use. Int. J. Occup. Environ. Health. 12, 260–267 (2006).
Turner, M. C. et al. Research recommendations for selected IARC-classified agents: impact and lessons learned. Environ. Health Perspect. 131, 105001 (2023).
Berrington de González, A. et al. Advisory Group recommendations on priorities for the IARC Monographs. Lancet Oncol. 25, 546–548 (2024).
Remigio, R. V. et al. An updated evaluation of atrazine-cancer incidence associations among pesticide applicators in the Agricultural Health Study Cohort. Environ. Health Perspect. 132, 27010 (2024).
Raghupathi, V. & Raghupathi, W. The influence of education on health: an empirical assessment of OECD countries for the period 1995–2015. Arch. Public Health 78, 20 (2020).
McNamara, C. L. et al. The socioeconomic distribution of non-communicable diseases in Europe: findings from the European Social Survey (2014) special module on the social determinants of health. Eur. J. Public Health 27, 22–26 (2017).
The Union for International Cancer Control (UICC). World cancer day 2020 international public opinion survey on cancer 2020. UICC https://www.uicc.org/resources/world-cancer-day-2020-international-public-opinion-survey-cancer-2020 (2022).
Perea, J. et al. Age at onset should be a major criterion for subclassification of colorectal cancer. J. Mol. Diagn. 16, 116–126 (2014).
Mounir, M. et al. New functionalities in the TCGAbiolinks package for the study and integration of cancer data from GDC and GTEx. PLoS Comput. Biol. 15, e1006701 (2019).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Zhou, W., Triche, T. J., Laird, P. W. & Shen, H. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nucleic Acids Res. 46, e123 (2018).
Hastie, T. Tibshirani, R. Narasimhan, B. & Chu, G. impute: impute: imputation for microarray data. R package v.1.84.0 https://bioconductor.org/packages/impute (2025).
Lin, S. M., Du, P., Huber, W. & Kibbe, W. A. Model-based variance-stabilizing transformation for Illumina microarray data. Nucleic Acids Res. 36, e11 (2008).
Chen, Y. et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 8, 203–209 (2013).
Ma, J. et al. Whole blood DNA methylation signatures of diet are associated with cardiovascular disease risk factors and all-cause mortality. Circ. Genom. Precis. Med. 13, e002766 (2020).
Liu, C. et al. A DNA methylation biomarker of alcohol consumption. Mol. Psychiatry 23, 422–433 (2018).
Madden, R. A. et al. Birth weight associations with DNA methylation differences in an adult population. Epigenetics 16, 783–796 (2021).
Do, W. L. et al. Epigenome-wide meta-analysis of BMI in nine cohorts: Examining the utility of epigenetically predicted BMI. Am. J. Hum. Genet. 110, 273–283 (2023).
Karabegović, I. et al. Epigenome-wide association meta-analysis of DNA methylation with coffee and tea consumption. Nat. Commun. 12, 2830 (2021).
van Dongen, J. et al. DNA methylation signatures of educational attainment. NPJ Sci. Learn. 3, 7 (2018).
Koh, I.-U. et al. Obesity susceptible novel DNA methylation marker on regulatory region of inflammation gene: results from the Korea Epigenome Study (KES). BMJ Open Diabetes Res Care. 8, e001338 (2020).
Joehanes, R. et al. Epigenetic signatures of cigarette smoking. Circ. Cardiovasc. Genet. 9, 436–447 (2016).
de, F. C. et al. Long-term air pollution exposure, genome-wide DNA methylation and lung function in the LifeLines Cohort Study. Environ. Health Perspect. 126, 027004 (2018).
Gondalia, R. et al. Methylome-wide association study provides evidence of particulate matter air pollution-associated DNA methylation. Environ. Int. 132, 104723 (2019).
Curtis, S. W. et al. Genome-wide DNA methylation differences and polychlorinated biphenyl (PCB) exposure in a US population. Epigenetics 16, 338–352 (2021).
Hoang, T. T. et al. Epigenome-wide DNA methylation and pesticide use in the agricultural lung health study. Environ. Health Perspect. 129, 97008 (2021).
Markunas, C. A. et al. Epigenome-wide analysis uncovers a blood-based DNA methylation biomarker of lifetime cannabis use. Am. J. Med. Genet. B Neuropsychiatr. Genet. 186, 173–182 (2021).
Tsaprouni, L. G. et al. Cigarette smoking reduces DNA methylation levels at multiple genomic loci but the effect is partially reversible upon cessation. Epigenetics 9, 1382–1396 (2014).
Zannas, A. S. et al. Epigenetic upregulation of FKBP5 by aging and stress contributes to NF-κB-driven inflammation and cardiovascular risk. Proc. Natl Acad. Sci. USA 116, 11370–11379 (2019).
Goobie, G. C. et al. Epigenome-wide analysis identifies pollution-sensitive loci in fibrotic interstitial lung disease. Am. J. Respir. Crit. Care Med. 211, 1835–1845 (2025).
Das, S. et al. Molecular and cell phenotype programs in oral epithelial cells directed by co-exposure to arsenic and smokeless tobacco. Biofactors 51, e70011 (2025).
Li, Q. S., Morrison, R. L., Turecki, G. & Drevets, W. C. Meta-analysis of epigenome-wide association studies of major depressive disorder. Sci. Rep. 12, 18361 (2022).
Rojas, D. et al. Prenatal arsenic exposure and the epigenome: identifying sites of 5-methylcytosine alterations that predict functional changes in gene expression in newborn cord blood and subsequent birth outcomes. Toxicol. Sci. 143, 97–106 (2015).
Green, B. B. et al. Epigenome-wide assessment of DNA methylation in the placenta and arsenic exposure in the New Hampshire Birth Cohort Study (USA). Environ. Health Perspect. 124, 1253–1260 (2016).
Kashima, K. et al. Identification of epigenetic memory candidates associated with gestational age at birth through analysis of methylome and transcriptional data. Sci. Rep. 11, 3381 (2021).
Paquette, A. G. et al. Regions of variable DNA methylation in human placenta associated with newborn neurobehavior. Epigenetics 11, 603–613 (2016).
Quinn, E. B., Hsiao, C. J., Maisha, F. M. & Mulligan, C. J. Prenatal maternal stress is associated with site-specific and age acceleration changes in maternal and newborn DNA methylation. Epigenetics 18, 2222473 (2023).
Cardenas, A. et al. The nasal methylome as a biomarker of asthma and airway inflammation in children. Nat. Commun. 10, 3095 (2019).
Kuan, P. F. et al. An epigenome-wide DNA methylation study of PTSD and depression in World Trade Center responders. Transl. Psychiatry 7, e1158 (2017).
Shen, X. et al. DNA methylome-wide association study of genetic risk for depression implicates antigen processing and immune responses. Genome Med. 14, 36 (2022).
Hannon, E. et al. Variable DNA methylation in neonates mediates the association between prenatal smoking and birth weight. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180120 (2019).
Küpers, L. K. et al. Meta-analysis of epigenome-wide association studies in neonates reveals widespread differential DNA methylation associated with birthweight. Nat. Commun. 10, 1893 (2019).
Choudhary, P. et al. Maternal educational attainment in pregnancy and epigenome-wide DNA methylation changes in the offspring from birth until adolescence. Mol. Psychiatry 29, 348–358 (2024).
Sharp, G. C. et al. Paternal body mass index and offspring DNA methylation: findings from the PACE consortium. Int. J. Epidemiol. 50, 1297–1315 (2021).
Sharp, G. C. et al. Maternal BMI at the start of pregnancy and offspring epigenome-wide DNA methylation: findings from the pregnancy and childhood epigenetics (PACE) consortium. Hum. Mol. Genet. 26, 4067–4085 (2017).
Viechtbauer, W. Conducting meta-analyses in R with the metafor package. J. Stat. Softw. 36, 1–48 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Method. 57, 289–300 (1995).
Tsai, H.-H. D. et al. Informing hazard identification and risk characterization of environmental chemicals by combining transcriptomic and functional data from human-induced pluripotent stem-cell-derived cardiomyocytes. Chem. Res. Toxicol. 37, 1428–1444 (2024).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 14, 7 (2013).
Mayakonda, A., Lin, D.-C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Korotkevich G, et al. Fast gene set enrichment analysis. Preprint at BioRxiv https://doi.org/10.1101/060012 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
SEER*Stat v.8.4.1 (National Cancer Institute, 2025).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmertest package: tests in linear mixed effects models. J. Stat. Softw. 82, 1–26 (2017).
R Core Team. R: a Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019).
Maas, S. C. E. et al. Epigenetic fingerprints link early-onset colon and rectal cancer to pesticide exposure. GitHub github.com/CancerCompBioLab/EOCRCexposome (2026).
Acknowledgements
First and foremost, we thank the participants and their families who took part in the studies analyzed. We are grateful to the investigators and data management teams who recruited the participants and to the pathologists who collected the samples. We thank J. Carmona and members of the Cancer Computational Biology Group for the helpful discussions. The results shown here are based, in part, on data generated by the TCGA Research Network (https://www.cancer.gov/tcga) and the SEER Program (www.seer.cancer.gov) SEER*Stat Database: Incidence—SEER Plus Research Data, 8 Registries, November 2021 submission (1975–2019)—Linked To County Attributes based on the November 2021 submission. This research was funded by the Instituto de Salud Carlos III (FORT23/00034), cofinanced by the European Regional Development Fund/European Social Fund ‘A way to make Europe’/‘Investing in your future.’ Grants CMS2022-135428, RYC2019-026576-I (to J.A.S.), JDC2022-048829-I (to S.C.E.M.) and FPU22/03520 (to M.B.E.) funded by MICIU/AEI/10.13039/501100011033 and, as appropriate, by ‘ESF Investing in your future,’ by ESF+ or by European Union NextGenerationEU/PRTR. Grant PID2020-115097RA-I00 funded by MICIU/AEI/ 10.13039/501100011033 (to J.A.S.) and, as appropriate, by ‘ERDF A way of making Europe,’ by ERDF/EU, by the European Union or by the European Union NextGenerationEU/PRTR and Ayuda Postdoctoral AECC POSTD258824MAAS (to S.C.E.M.). The research leading to these results has received funding from la Caixa Foundation. This work was supported by the VHIO-CRIS Program for Precision Oncology and Digitalization from Fundación CRIS Contra el Cáncer. VHIO would like to acknowledge the State Agency for Research (Agencia Estatal de Investigación) for the financial support as a Center of Excellence Severo Ochoa (CEX2020-001024-S/AEI/10.13039/501100011033), the Cellex Foundation for providing research facilities and equipment and the CERCA Program from the Generalitat de Catalunya for their support on this research. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Ethics declarations
Competing interests
I.B. reports accommodation and travel expenses from Amgen, Merck, Sanofi and Servier; and personal speaker honoraria from AstraZeneca and Amgen. E.E. has received personal honoraria from Agenus, Amgen, Bayer, Boehringer Ingelheim, Bristol Myers Squibb, Cure Teq AG, GlaxoSmithKline, Hoffman La Roche, Janssen, Johnson&Johnson, Lilly, Medscape, Merck Serono, MSD, Nordic Group BV, Novartis, Organon, Pfizer, Pierre Fabre, Repare Therapeutics Inc., RIN Institute Inc., Rottapharm Biotech, Sanofi, Seagen International GmbH, Servier and Takeda. J.T. reports personal financial interest in form of scientific consultancy role for Accent Therapeutics, Alentis Therapeutics, AstraZeneca, Boehringer Ingelheim, Bristol Myers Squibb, Carina Biotech, Cartography Biosciences, Chugai, Daiichi Sankyo, F. Hoffmann-La Roche, Genentech, Johnson & Johnson/Janssen, Larkspur Biosciences, Lilly, Marengo Therapeutics, Menarini, Merus, MSD, Novartis, One-carbon Therapeutics, Ono Pharma USA, Peptomyc, Pfizer, Pierre Fabre, Quantro Therapeutics, Scandion Oncology, Scorpion Therapeutics, Servier, Sotio Biotech, Syntelios AG, Taiho, Takeda Oncology, Theriva Biologics and Tolremo Therapeutics; and stocks from Alentis Therapeutics, Oniria Therapeutics, 1TRIALSP and Pangaea Oncology. The other authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Ulrike Harjes, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The relationship between directly measured exposome traits and their respective methylation risk scores.
The plots show the distribution of measured exposures compared with the constructed MRSs for A. the alcohol MRS-GW in TCGA liver hepatocellular carcinoma tumors, comparing patients with (N = 122) and without (N = 64) alcoholic liver disease, B. for the smoking-Maas MRS in blood samples of non-cancer patients comparing current (N = 22), former (N = 263), and never (N = 179) smokers, and C. in TCGA tumor samples, comparing current (N = 505), former (N = 838), and never (N = 453) smokers, with D. further sub-categorization of former smokers based on >15 years (N = 300) vs. ≤15 years (N = 538) since smoking cessation in the tumor samples, including cervical squamous cell carcinoma and endocervical adenocarcinoma, esophageal carcinoma, head and neck squamous cell carcinoma, lung adenocarcinoma, lung squamous cell carcinoma, and pancreatic adenocarcinoma (Supplementary Table S4). Distribution of E. the cannabis MRS-P1E5 in whole blood samples comparing cannabis dependent (N = 30) with non-dependents (N = 242) and F. of the PM2.5 MRS-P1E5 comparing a high 5-year median PM2.5 exposure (The University of Pittsburgh; N = 308; 12.1 µg/m3) to a lower median exposure (University of British Columbia; N = 171; 5.1 µg/m3). The differences between groups were evaluated using the Wilcoxon rank-sum test. The y-axes reflect the exposure MRS and the CpG threshold cutoff, while the x-axes show the measured exposure and the dataset in which the MRS was validated. Box plots show the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers.
Extended Data Fig. 2 The relationship between measured exposures and their respective methylation risk scores.
The plots show the distribution of measured exposures compared with the constructed MRSs for A. asthma MRS-GW in 395 asthma patients and 246 non-asthmatic controls, for B. prenatal exposure to arsenic MRS-P1E5 in 6 exposed vs. 6 non-exposed htert-immortalized normal oral keratinocytes (NOK) cells, for C. major depressive disorder MRS-P1E5, and D. a polygenic risk score (PRS) for depression MRS-P1E5 in 361 depressive patients and 68 controls. The differences between groups were evaluated using the Wilcoxon rank-sum test. The y-axes reflect the exposure MRS, the CpG threshold cutoff, and the PubMed ID of the original EWAS, while the x-axes show the measured exposure and the dataset in which the MRS was validated. Box plots show the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers.
Extended Data Fig. 3 The relationship between prenatal exposome exposures and their corresponding methylation risk scores across tissues collected at birth.
The plots show the distribution of measured prenatal exposure compared with the constructed MRSs for arsenic exposure A. MRS-GW in cord blood (N = 38) and B. MRS-P1E5 in the placenta (N = 343), C-D. for two maternal BMI MRSs-P1E5 in cord blood (N = 110), E-H. for two birthweight MRS-P1E5 in E-F. cord blood (N = 110) and G-H. venous blood (N = 176), I-J. for the maternal education MRS-P1E5 in two placental cohorts (N = 319 and 334), and the distribution of gestational age MRS-GW in K. venous blood (N = 176) and L. cord blood (N = 110). The correlations were assessed using Pearson’s correlation, and the differences between groups were evaluated using the Wilcoxon rank-sum test. The y-axes reflect the exposure MRS, the CpG threshold cutoff, and the PubMed ID of the original EWAS, while the x-axes show the measured exposure and the dataset in which the MRS was validated. Box plots show the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers.
Extended Data Fig. 4 Schematic overview to determine directionality of the methylation risk scores.
For CpG sites that are positively associated with the exposome trait in the original epigenome-wide association study (EWAS), a higher methylation level reflects a higher exposure (orange), while for negatively associated CpG sites, a lower methylation level reflects a higher exposure (blue). When sorting the patients in TCGA-COAD based on their methylation risk score (MRS), the heatmaps presented in Figs. 2 and 3 identified a pattern in which the EWAS-positive CpG sites had higher methylation levels and EWAS-negative CpG sites had lower methylation levels, correlating with a high MRS. Consequently, we deduced that a higher MRS equals a higher exposure to the exposome trait.
Extended Data Fig. 5 Late-onset colon cancer patients are more likely to be obese than early-onset patients.
A subset of 96 patients within the TCGA-COAD cohort had physical metrics available. A. Among 24 early-onset patients, 4 were classified as obese (BMI > 30 kg/m2), compared to 18 out of 72 late-onset patients. This provided a relative risk (RR) of 0.67 (95% confidence interval: 0.26-1.76) for obesity in early-onset colorectal cancer patients. B. The boxplots show the methylation risk score distribution among these 96 patients, including the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers. CI; Confidence Interval, RR; relative risk.
Extended Data Fig. 6 Exposome-related methylation risk scores show differences between early-onset and late-onset colon and rectal cancer patients.
Summary overview of the association between exposome-related methylation risk scores (MRS) comparing early-onset (EO; age <50) to late-onset (reference group) cases (LO; age ≥70) by employing logistic regression analyses adjusted for sex. The left column shows the results obtained in the discovery phase, conducted in TCGA-COAD (EO = 31 vs LO = 100). The rightmost panel shows the results obtained in the replication phase, comprising the meta-analysis of nine cohorts (EMTAB3027, EMTAB7036, GSE199057, GSE131013, GSE42752, TCGA-READ, GSE39958, GSE101764, and GSE77954), including 83 EOCRC and 272 LOCRC patients. Separate meta-analyses were performed using the datasets generated with the Illumina Infinium Human Methylation 450 K BeadChip array, excluding GSE199057, and for exploratory analysis in a tissue-specific analysis for rectal cancer samples (TCGA-READ and GSE39958) and colon cancer samples (EMTAB7036, GSE199057, GSE131013, and GSE42752). The marker selection thresholds indicate the significance threshold in the original EWAS used for CpG selection, namely P < 1.2×10−7 (GW), P < 1.0×10−5 (P1E5), and false discovery rates (FDR) of <0.1 (F01), <0.05 (F005), and <0.01 (F001). The presence of CpGs across the five marker selection thresholds varies, with unavailable data indicated in white as N.A., while non-significant findings are represented in light grey. 2,4-D; 2,4-Dichlorophenoxyacetic acid, AHEI; Alternative Healthy Eating Index, BMI; body mass index, DDT; Dichlorodiphenyltrichloroethane, MDS; Mediterranean Diet Score, NO2; nitrogen dioxide, PCB; polychlorinated biphenyls, PM2.5, and PM2.5-10; particulate matter <2.5, and between 2.5 and 10 micrometers (µm) in diameter, smoking-Maas; CpGs from smoking inference model without additional threshold.
Extended Data Fig. 7 Picloram MRS-GW distribution in clinical characteristics.
The boxplots show the distribution of the Picloram MRS-GW across A. sex, B. MSI status, C. Consensus Molecular Subtypes (CMS), and D. tumor purity set at ≥0.7 for high and <0.7 for low. The left panels show the complete data sets, while the right panels show the distribution across the age at diagnosis categories, including early- (EOCRC; aged <50), intermediate- (IOCRC; aged 50 to 69 years), and late-onset (LOCRC; aged ≥70 years). Box plots show the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers.
Extended Data Fig. 8 Permutation results for CpG sites and patient categorization.
The violin plots show the results from the permutation tests. A. The stability of the CpGs in each MRS was tested using 10,000 permutations. The red asterisks reflect the p-values obtained using the CpGs included in our MRSs. B. The patient categorization is tested using 1,000 permutations. The red asterisks reflect the p-values categorization using age at diagnosis for each MRS. Box plots show the median (center line), interquartile range (box), and whiskers extending to the most extreme values within 1.5×IQR; points beyond represent outliers.
Extended Data Fig. 9 Molecular differences between high and low Picloram MRS-GW in TCGA-COAD.
Tumors were stratified into tertiles according to the picloram MRS-GW, and the lowest (N = 76) and highest (N = 76) tertiles were compared using Fisher’s Exact Test to identify differentially mutated genes (P < 0.05). A. Percentage of tumors harboring mutations in each gene in the lowest and highest tertiles in TCGA-COAD. B. Distribution of mutations across all tumors, ordered by picloram MRS-GW.
Extended Data Fig. 10 Mutational signature accuracy and the relationship of SBS1 score distribution with age and picloram MRS-GW.
A. The plot shows the distribution of cosine similarity (accuracy) across MSS tumor samples. The histogram (orange bars) shows the counts by cosine similarity score, while the overlaid points represent tumor mutation burden for each sample. The dashed vertical line indicates the mean cosine similarity threshold. B. The distribution of the single-base substitution signature 1 (SBS1) score shows the relation with chronological age in microsatellite stable (MSS) tumors (blue), while microsatellite instable-high (MSI-H) tumors (orange) show a distinct pattern that differs from MSS tumors and chronological age. C. The SBS1 score distribution in patient categorization based on age at diagnosis, including early- (aged <50; N = 25), intermediate- (aged 50 to 69 years; N = 76), and late-onset (aged ≥70 years; N = 72) cases. Next, patients were categorized based on the acquired mutations belonging to SBS1 as young tumors ( < 60) and old tumors ( ≥ 60). The boxplot shows D. the picloram-MRS distribution and E. the chronological age distribution across the SBS1-based categorization. EOCRC; early-onset CRC, IOCRC; intermediate-onset, LOCRC; late-onset CRC, SBS1; single-base substitution signature 1.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Maas, S.C.E., Baraibar, I., Lemler, L. et al. Epigenetic fingerprints link early-onset colon and rectal cancer to pesticide exposure. Nat Med (2026). https://doi.org/10.1038/s41591-026-04342-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41591-026-04342-5