Main
At present, 537 million adults worldwide are living with diabetes, a figure that is estimated to increase to 643 million by 2030. Approximately 10% of people with diabetes have type 1 diabetes (T1D), and around 90% have type 2 diabetes (T2D)4. The rise in T2D is driven mainly by lifestyle factors5. In a healthy individual, insulin—a hormone secreted by pancreatic β-cells—helps to regulate blood glucose levels by facilitating the uptake of glucose from the blood into cells (including muscles, adipose and liver). In addition, incretin hormones, such as glucagon-like peptide 1 (GLP1) and gastric inhibitory polypeptide (GIP), can increase the secretion of insulin from pancreatic β-cells, leading to improved glycaemic control6. The fundamental problem in diabetes is the inability of the body to regulate blood glucose properly owing to absolute or relative insulin deficiency. In T1D, the body’s immune system mistakenly attacks and destroys the pancreatic β-cells, resulting in absolute insulin deficiency and high blood glucose7. By contrast, in most cases of T2D, the body becomes insulin resistant, meaning that higher amounts of insulin have to be produced by the pancreatic β-cells to achieve the same glucose-lowering effect. With time, the pancreatic β-cells can become unable to produce enough insulin to compensate for IR, leading to relative insulin deficiency and increased blood glucose levels. Figure 1a illustrates the complex nature of T2D and the intricate relationships between lifestyle choices, genetics and various metabolic subphenotypes and physiological processes that are involved in the development of the disease. Long-term complications of diabetes include damage to various organs and tissues over time, such as diabetic retinopathy, nephropathy and neuropathy8.
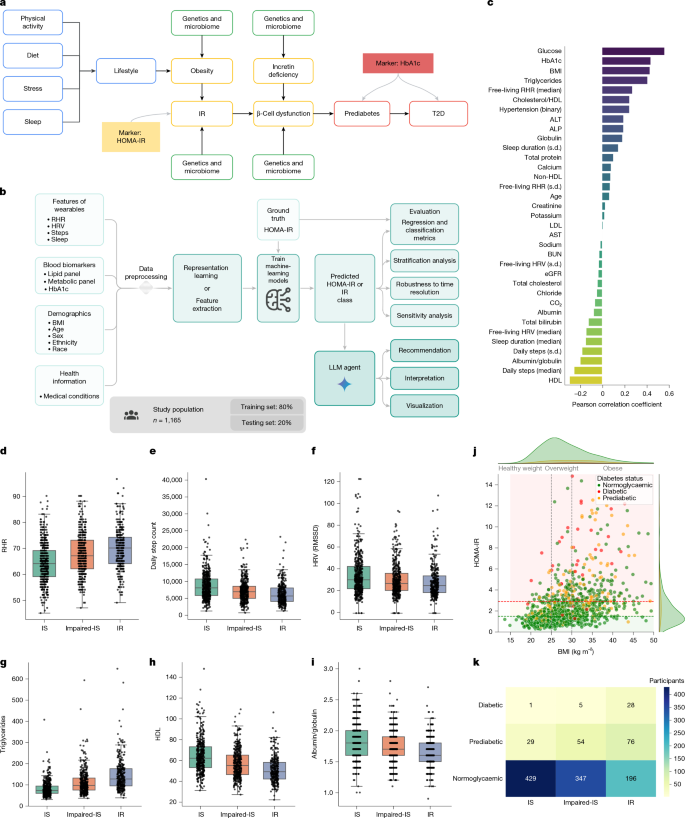
a, Overview of physiological factors and associated lifestyle factors leading to IR, prediabetes and diabetes. b, Our proposed modelling pipeline for predicting HOMA-IR and interpreting the results with the insulin resistance literacy and understanding agent (IR agent). c, Correlation of blood biomarkers and lifestyle features (continuous values) with HOMA-IR. d–f, Distribution of the top three features of wearables that are highly correlated with HOMA-IR (RHR (d), daily step counts (e) and HRV (f)) for stratified insulin sensitivity groups (IS, impaired-IS and IR). RMSSD, root mean square of successive differences. g–i, Distribution of the top three highly correlated blood biomarkers (triglycerides (g), HDL cholesterol (h) and albumin/globulin ratio (i)) for stratified insulin sensitivity groups. In the box plots in d–i, the centre line indicates the median, the bounds of box represent the 25th and 75th percentiles and the whiskers extend to 1.5 times the interquartile range. j, Scatter plot of BMI and HOMA-IR values, showing the relationship between higher BMI values and IR (measured through HOMA-IR). k, Confusion matrix showing the number of participants in each combination of IR status and diabetes status.
The prevalence of IR in the general population is estimated to be between 20% and 40%, with variation observed across ethnic groups, age brackets, lifestyle and the presence of comorbidities1. IR prevalence in T2D is 83.9% (ref. 9). Factors that contribute to IR include excess body weight (particularly visceral fat), physical inactivity and genetic predisposition. Chronic IR puts the person at considerable risk of prediabetes and overt T2D. Moreover, IR is strongly associated with the risk of metabolic-dysfunction-associated steatotic liver disease (MASLD) and cardiovascular disease (CVD).
Identifying IR early can guide several focused lifestyle interventions, such as weight loss, regular exercise and a healthy dietary pattern, that can substantially improve or even reverse IR. Although most individuals can certainly benefit from most types of physical activity and healthy diets, specific interventions have been scientifically shown to prevent and treat IR. For lifestyle interventions, resistance training10,11, aerobic training10, calorie-restricted diets12 and low-fat diets13 are all valuable in reducing IR. On the therapeutic side, thiazolidinediones and metformin medications have frequently been shown to reduce IR14,15. Studies have shown that incretin-hormone agonists, such as GLP1 and GIP, act as sensitizers and improve IR16,17.
Several methods are available for assessing IR, but are not implemented routinely, meaning that opportunities for early intervention are often missed. Instead, a focus on snapshots of glucose levels, fasting glucose, HbA1c or glucose levels after a two-hour oral glucose tolerance test (OGTT) represents the typical screening approach, and can be insensitive to those in the early stages of IR. The gold-standard test for IR is the hyperinsulinaemic euglycaemic clamp2, which is performed in research facilities only, and is expensive and time consuming. Homeostatic model assessment of IR (HOMA-IR) is a more affordable and faster alternative, but it requires a clinical laboratory visit3. Glucotyping, which is a framework that analyses glucose time-series data from continuous glucose monitoring (CGM), is a recent method to detect IR that can be done at home, but it requires further validation studies18,19. Physiological signals derived from smartwatches could conceivably help to predict IR, because it has been shown that higher resting heart rate (RHR) and lower heart-rate variability (HRV) are associated with IR20,21,22,23.
In this study, we present a method for predicting IR using signals derived from a consumer smartwatch, demographics and routinely measured blood biomarkers. This method has the potential to be scaled to millions of people, and to enable widespread identification of IR. We assembled a large cohort (n = 1,165) with a combined set of data from wearable devices, together with demographics and blood biomarkers, and a ground-truth measure of IR (HOMA-IR). We performed a comprehensive analysis of our model, including interoperability, stratification and robustness analyses, to quantify its generalizability and scalability. Furthermore, we developed a large language model (LLM) agent that uses the output of the IR model, along with participant lifestyle and blood biomarker data, to provide safe, holistic insights into individual metabolic health and diabetes risk, and to offer personalized recommendations and illustrative explanations.
Study design and cohort characteristics
We designed the Wearables for Metabolic Health (WEAR-ME) study and recruited adults from the USA to take part. Participants provided informed consent, and the study was approved by Advarra (Institutional Review Board (IRB) no. Pro00074093). The study was conducted remotely using the Google Health Studies (GHS) application (Methods). In this instance, GHS was configured to enable the collection of data from Fitbit and Google Pixel watch devices (collectively referred to throughout as wearables), completion of questionnaires and ordering of blood tests with Quest Diagnostics.
We used HOMA-IR, calculated as HOMA-IR = (fasting insulin (µU ml−1) × fasting glucose (mg dl−1))/405, as the ground truth for quantifying IR (Methods). The thresholds of HOMA-IR used to define IR vary widely in the literature, ranging from 2.5 to 3.5 for significant IR and 1 to 1.5 for insulin sensitivity1,24,25,26. The differences in thresholds are attributable mainly to differences in study populations (ethnicity, age and gender), and the specific factor used to define the threshold (either maximizing the sensitivity for predicting metabolic syndrome or using the 90th percentile of the study population). In this study, participants were classified as having IR (HOMA-IR > 2.9), insulin sensitivity (IS) (HOMA-IR < 1.5) or as having impaired insulin sensitivity (impaired-IS) (1.5 ≤ HOMA-IR ≤ 2.9). Extended Data Table 1 summarizes the characteristics of our cohort. A total of 1,165 individuals (459 with IS, 406 with impaired-IS, 300 with IR) with high-quality data (Supplementary Fig. 1) were included in the development of the IR model. Supplementary Fig. 2 shows the distribution of the seven different smartwatches and four trackers in the WEAR-ME cohort. Participants with IR had higher rates of diabetes, CVD, hyperlipidaemia and hypertension. Supplementary Table 1 summarizes all digital and blood biomarkers among the three groups.
Figure 1b illustrates the design of our deep learning framework for predicting IR. It takes various combinations of data from wearables, blood biomarkers, demographics and health information as input. Time-series data are preprocessed and summarized, and an embedded representation is extracted through a masked autoencoder (MAE) (Methods). This representation is fed into multiple tree-based models to predict continuous HOMA-IR values. Predicted IR classes (IR or non-IR) were obtained by thresholding on the predicted HOMA-IR using a HOMA-IR threshold of 2.9. We tested 25 combinations of input features (wearable features, demographics, fasting glucose, lipid panel, HbA1c, metabolic panel and hypertension status). Training and testing were done using fivefold cross-validation for direct regression models. Interpretability analysis was performed to assess the contributions of the features to the learned representation and the model performance (Supplementary Information). We performed a comprehensive evaluation of the predicted HOMA-IR and IR classes to assess the generalizability and scalability of the trained model. In addition, we performed a robustness and stability analysis to assess the variability of the model’s output for each individual in the dataset, on the basis of varying time intervals of the data from wearables (one week, two weeks and up to three months; Supplementary Information).
Association of IR with lifestyle and blood biomarkers
We calculated Pearson correlation coefficients between HOMA-IR and major lifestyle factors (RHR, HRV, step count and sleep duration), demographics, lipids, glucose, markers of kidney and liver function and key electrolytes (Supplementary Table 2). Figure 1c shows the significant positive correlations between HOMA-IR and fasting glucose (r = 0.57, P < 0.001), BMI (r = 0.43, P < 0.001; Supplementary Fig. 4), HbA1c (r = 0.45, P < 0.001), triglycerides (r = 0.40, P = < 0.001; Fig. 1g) and RHR (r = 0.27, P = < 0.001; Fig. 1d). Moreover, HOMA-IR showed a significant negative correlation with high-density lipoprotein (HDL) cholesterol (r = −0.30, P < 0.001; Fig. 1h), daily step count (r = −0.25, P = < 0.001; Fig. 1e), albumin/globulin ratio (r = −0.18, P = < 0.001; Fig. 1i) and HRV (r = −0.14, P < 0.001; Fig. 1f). This suggests that HOMA-IR can be inferred using readily available measures from wearables or blood biomarkers. Age, kidney markers (for example, creatinine, estimated glomerular filtration rate (eGFR) and blood urea nitrogen (BUN)) and electrolytes (for example, sodium, potassium and chloride) have a low Pearson correlation coefficient with HOMA-IR (|r| < 0.1). C-reactive protein (CRP) levels were significantly higher in the IR group than in the IS group (2.8 mg dl−1 versus 0.6 mg dl−1, P < 0.001). Analytes assessed in the standard complete blood count (for example, white blood cell, red blood cell, haemoglobin, haematocrit and so on) did not differ significantly in their effect size between the IR and IS groups (Supplementary Table 1). Figure 1j illustrates the relationship between obesity (measured by BMI) and IR, as assessed by HOMA-IR. A total of 205 out of 458 (45%) individuals with obesity (BMI > 30) are insulin resistant (HOMA-IR > 2.9). Only 22 out of 319 (6.9%) participants with a normal healthy weight (18.5 < BMI < 25) are insulin resistant. Figure 1k highlights the relationship between IR and diabetes. In the WEAR-ME study cohort, 33 out of 34 (97%) individuals with diabetes (HbA1c > 6.5%) are classified as having either IR or impaired-IS, whereas only one participant has IS. Notably, individuals may have IR without an apparent increase in HbA1c levels. A total of 196 out of 972 (20%) normoglycaemic participants have IR, representing those at high risk of developing diabetes. This underscores the importance of identifying these individuals early to enable personalized lifestyle interventions that could potentially reverse the course of T2D development. Supplementary Fig. 4 shows the pairwise correlation between HOMA-IR, wearable features, demographics and blood biomarkers.
IR prediction using wearables and blood biomarkers
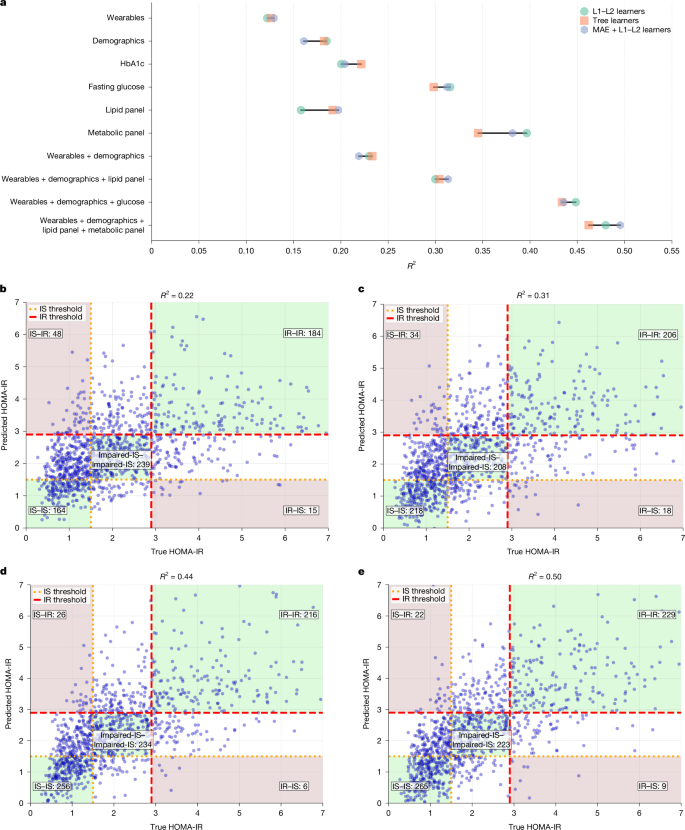
We trained multimodal models using various combinations of wearable features, demographics and blood biomarkers to predict IR. Specifically, we trained regression models to predict continuous HOMA-IR values, and then applied classification thresholds to determine IR status. Figure 2 illustrates the regression performance for selected sets of features, using a seven-day aggregation window for wearable features. Figure 2a shows R2 values for both direct regression (XGBoost with linear (L1–L2) and non-linear (tree) learners) and representation learning (MAE + XGBoost with linear learners (L1–L2)) (Methods). Our results show that incorporating data from wearables into models, as well as using demographic information and readily available blood biomarkers, significantly enhances prediction accuracy. Furthermore, our analyses show an increase in true positives (shaded light green) and a reduction in consequential false predictions (participants who are predicted to have IR, but have IS; shaded light brown area in Fig. 2b–e). Most notably, the addition of fasting glucose alone doubled the R2 value from 0.212 to 0.435 (Fig. 2d), increased the number of correctly identified individuals with IR by 17% (from 184 to 216) and reduced consequential false positives (participants who are identified as having IR, but have IS) by 46% (from 48 to 26). Meanwhile, our experiments showed that using fasting glucose alone is not sufficient (R2 = 0.31), highlighting the importance of other lifestyle factors in estimating HOMA-IR. The optimal model for predicting HOMA-IR combines data from wearables, demographics and readily available blood biomarkers (fasting glucose, lipid panel and metabolic panel) (R2 = 0.50; Fig. 2e).
a, Comparison of HOMA-IR regression across input feature sets and models. b–e, Scatter plots of predicted HOMA-IR values versus the true HOMA-IR models for selected feature sets: wearables and demographics (b), wearables, demographics and lipid panel (c), wearables, demographics and fasting glucose (d) and wearables, demographics, lipid panel and metabolic panel (e). Areas of concern for true positive and false negative are highlighted as light green and light brown, respectively.
Subsequently, we performed a rigorous evaluation of the capacity of the models to accurately classify IR, using the predicted HOMA-IR values with a threshold of 2.9. Extended Data Fig. 1a shows that a model based on wearables and demographics alone can predict IR with AUROC = 0.70, sensitivity = 0.60 and specificity = 0.80. Incorporating fasting glucose levels into this model resulted in a significant improvement in performance (AUROC = 0.78, sensitivity = 0.73, specificity = 0.84). A model that includes wearables, demographics, readily available blood biomarkers (lipid panel and metabolic panel) yielded values of AUROC = 0.80, sensitivity = 0.76 and specificity = 0.84. It is crucial to underscore that relying on demographics, wearables, fasting glucose or lipid panels in isolation does not yield adequate predictive power for IR (Extended Data Fig. 1a and Supplementary Information). A comprehensive summary of the experimental settings, performance benchmarks and ablation analyses is provided in Supplementary Tables 3–6. Supplementary Tables 7 and 8 report the statistical significances of the differences in AUROC between each pair of experiments, as determined by Wilcoxon rank-sum and McNemar’s test, respectively. To further elucidate the performance of the models under various predicted HOMA-IR thresholds, Extended Data Fig. 1b,c present the ROC and precision-recall curves, respectively, for the four feature sets that can be practically implemented from available data. Our findings show integrating data from wearables, demographics and readily accessible blood biomarkers significantly enhances our ability to predict IR, compared with relying on each data source in isolation. The interpretability of the learned representation and prediction models is shown in Extended Data Fig. 2, Supplementary Table 9 and Supplementary Information, along with a stratified performance analysis of IR prediction based on BMI and physical activity,
IR prediction with a wearable foundation model
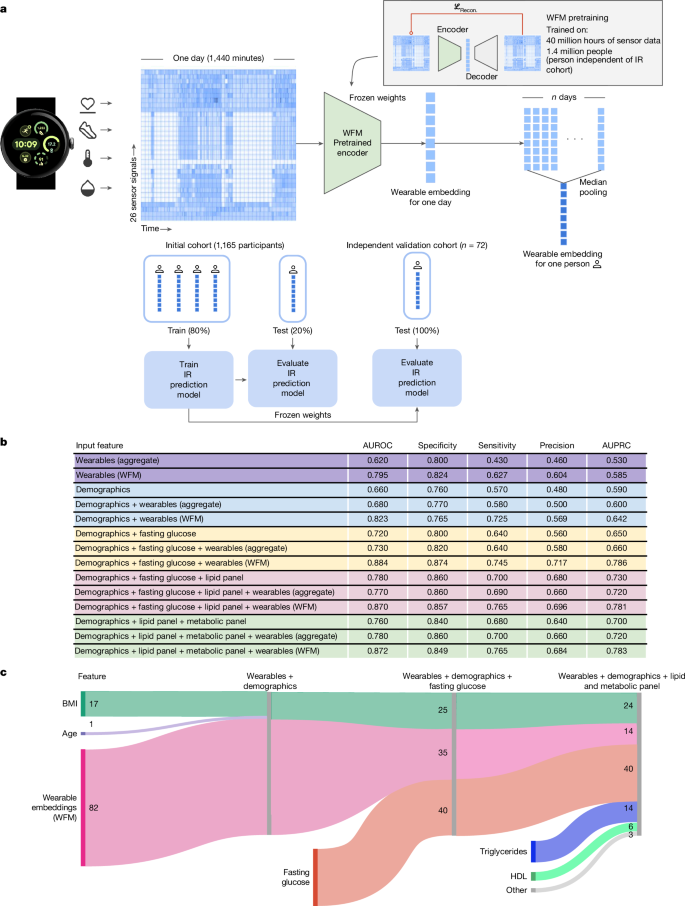
Subsequently, we took a holistic approach to modelling the wearable-device data, using a foundation model to learn robust high-dimensional feature representations of complex data. Foundation models have become a key tool in scientific analysis, because they provide a way to learn robust high-dimensional feature representations of complex data from unlabelled examples using pretraining pre-text tasks27,28. We investigated whether a foundation model trained on a large corpus of data from wearables could lead to an improvement in IR prediction. Our pretrained WFM extracted representations from wearables input data at one-minute resolution. The input window size was one day (1,440 min) by 26 signals (Methods). Given that each participant has many days of data from wearables, we used a median pooling to generate a single embedding per participant. These embeddings have dimensions of 384. To reduce the dimensionality, principal component analysis was fitted on the training set and the mapping applied to the test set. The first five principal components were used for the downstream task of IR prediction. We then fine-tuned a non-linear classification head on the five top principal components from the median-pooled, frozen wearable signal embeddings from the foundation model (Fig. 3a).
a, Schematic representation of the WFM pretraining and inference components. L Recon., reconstruction loss. b, Performance metrics quantifying the added value of the WFM in improving IR prediction. AUPRC, area under precision-recall curve. c, SHAP analysis quantifying the relative contribution of wearable embeddings from the WFM to prediction performance for various experimental settings.
Using feature embeddings from the WFM improved the predictive power of IR, compared with using aggregate wearable measures. In 80% (932 participants) and 20% (233 participants) of the initial WEAR-ME cohort, the training set and test set, respectively, a model integrating WFM-derived representations with demographics surpassed a demographics-only baseline (AUROC = 0.82 versus 0.66; Fig. 3b). Moreover, adding WFM representations to a model with demographics, fasting glucose and a lipid panel substantially improved predictive performance over an identical model without wearable-device data (AUROC = 0.87 versus 0.78; Supplementary Table 10). All performance metrics for the fivefold cross-validation experiments are described in Supplementary Table 11.
Furthermore, the Shapley additive explanations (SHAP) feature importance shows that, for models based on the WFM embeddings, the contribution of the wearable-device data is higher (82% for the WFM; Fig. 3c) than it is with a conventional machine-learning model (43%; Extended Data Fig. 2a). These results show that the detailed embedding of the WFM captures the complex dynamics and interplay of daily activity, sleep and physiological rhythms that are crucial to IR but missed by simple aggregates.
Validation in an independent cohort
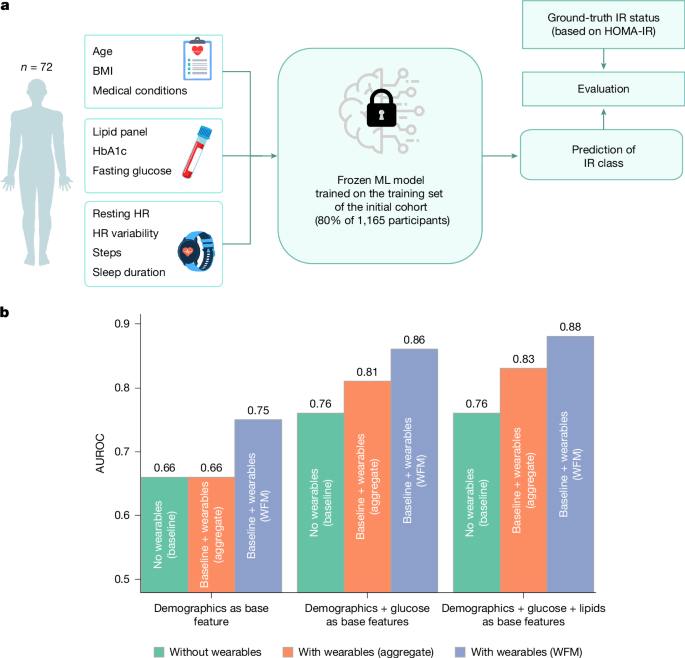
To validate the generalizability of the proposed IR models to unseen data, we evaluated the performance of our trained IR prediction model on an independent validation cohort. The study protocol was approved by WCG (IRB no. 1371945) (Methods). Data collected throughout the study included anthropometric measurements (for example, BMI, waist circumference and skin tone), blood biomarkers (for example, HbA1c, fasting glucose and lipid panel), wearable-device data (from a Fitbit Charge 6, measuring heart rate, RHR, HRV, sleep duration and step count), ground-truth IR (HOMA-IR) and health and lifestyle questionnaires (Fig. 4a). To start with, 144 individuals were enrolled, among whom 127 individuals had complete wearable-device data and 82 individuals had complete physiological biomarker data acquired during an in-person visit at the end of the study (Methods). Ultimately, 72 individuals had both complete wearable-device data and complete physiological biomarker data, and this group was used to validate the generalizability of our IR prediction models. The cohort members had an average age of 44.5 years and BMI of 30.6 kg m2, and mixed ethnicities (Extended Data Table 2). Similar to the initial cohort (n = 1,165), HOMA-IR was used as the ground-truth measure of IR in the independent validation study. Using HOMA-IR cut-off values of 2.9 and 1.5, the validation cohort was divided into three groups: IS (33 people), impaired-IS (20 people) and IR (19 people).
a, Overview of the study for the independent validation cohort. b, Performance of the IR classification on the independent validation cohort based on various experimental settings, without wearables and with wearables (aggregate and with WFM). We validated all of our trained models on this cohort except models with a complete metabolic panel (CMP), because the independent validation cohort did not include a CMP in the blood tests.
For the validation experiments, we selected pretrained IR prediction models that had been trained on 80% of the initial cohort (WEAR-ME) and froze their weights (Supplementary Table 10). These included models with and without wearable-device data, in which data from wearables were represented either as simple aggregate measures (average RHR, HRV, step counts and sleep duration) or through the WFM. This approach allowed us to directly assess the added value of including wearable-device data. We then applied these models with frozen weights to the independent validation cohort. The results provide further evidence that data from wearables provide considerable added value in predicting IR, even when the model is applied to previously unseen data (Fig. 4b). A model integrating WFM-derived representations with demographics surpassed a demographics-only baseline (AUROC = 0.75 versus 0.66). Furthermore, adding WFM representations to an optimal model that included demographics, fasting glucose and a lipid panel substantially improved predictive performance over an identical model without data from wearables (AUROC = 0.88 versus 0.76). Supplementary Table 12 shows all evaluation metrics on the independent validation study.
IR literacy and understanding agent
The ability to detect IR from wearables and routine blood biomarkers before the onset of T2D (characterized by HbA1c > 6.5%) raises the possibility of using an LLM agent connected to users’ wearables and personal health records to notify people that they have an increased risk of developing diabetes. It could also incorporate the inferred user’s IR class interactively when they ask general queries related to metabolic health. To show the potential of such a system for answering metabolically relevant questions, we set out to design a reasoning agent (Methods), illustrated in Fig. 5a. Our proposed agent, called the insulin resistance literacy and understanding agent (IR agent) uses a reason and act (ReAct) framework that is built on top of an LLM—in our case, Gemini 2.0 Flash. Our agent combines the language understanding of an LLM with the ability to perform actions, such as searching the web for up-to-date information, accessing specialized tools like a calculator and using our IR prediction models. This allows the IR agent to dynamically plan its response to a user’s query about their metabolic health, grounding its answers in real-world data and verifiable calculations, rather than relying solely on the LLM’s pre-existing knowledge. After receiving a user query, a data frame of the user’s health data, along with the query itself, are provided to the agent within its context window, enabling the agent to tailor its reasoning and actions to the individual’s specific health profile.
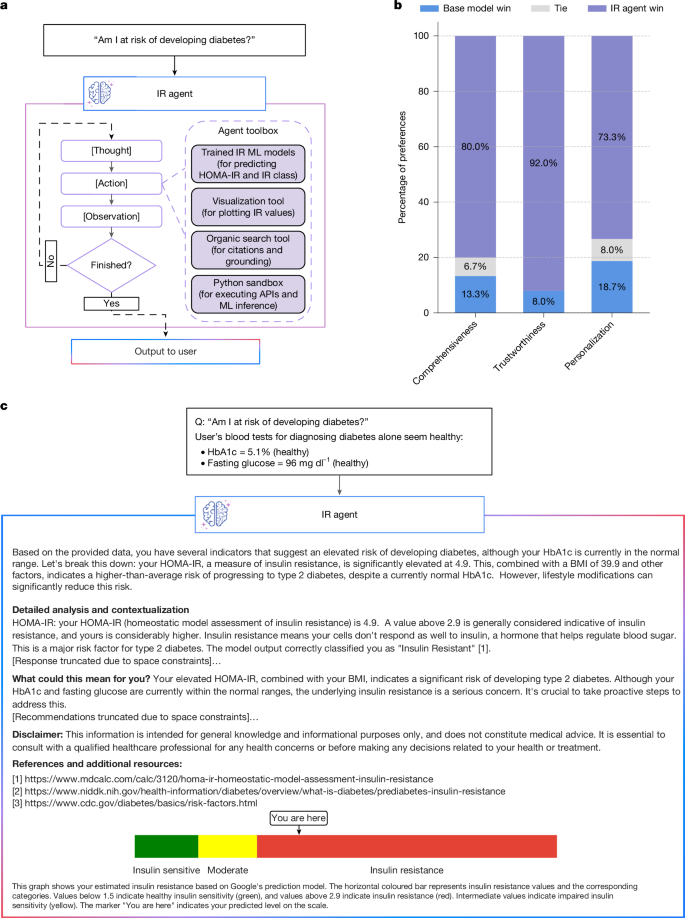
a, Illustration of the proposed IR agent. API, application programming interface; ML, machine learning. b, Win rate of our IR agent against the base model as evaluated by endocrinologists in side-by-side comparisons. c, Example of a metabolically relevant question paired with data from a real study participant, and the corresponding IR agent output.
We evaluated the efficacy and clinical relevance of the IR agent through a comprehensive assessment by five external, board-certified endocrinologists. This evaluation focused on two key aspects: (1) the added value of incorporating predicted IR information into an LLM response and (2) the absolute accuracy and clinical safety of the IR agent’s outputs (Methods). For the added-value assessment, endocrinologists performed blinded side-by-side comparisons of responses generated by the IR agent (with IR information) and a base LLM (Gemini 2.0 Flash, with access to the same user data but without IR prediction). These comparisons, using queries from five representative study participants with diverse metabolic profiles (Supplementary Table 13), revealed a strong preference for the IR agent across all evaluated dimensions (Fig. 5b and Supplementary Table 14). Specifically, the IR agent was rated as more comprehensive (80% preference), trustworthy (92% preference) and personalized (73.3% preference), compared with the base LLM. Figure 5c shows an example of a query and the response from the IR agent.
For the absolute-accuracy assessment, endocrinologists evaluated the responses of the IR agent across four dimensions: factuality; data referencing and interpretation; safety; and grounding (citation validity). The IR agent showed high factuality (79% of responses deemed completely factually accurate) and safety (96% of responses considered safe) (Supplementary Fig. 8). A detailed inferred analysis of the data referencing and interpretation component revealed that our agent was able to accurately reference and interpret HOMA-IR values (100% and 96%, respectively) and demographic data (96% for both referencing and interpretation). Although the referencing of wearables and blood biomarkers was consistently accurate, the interpretation accuracy was lower for these data types (79% and 59%, respectively; see Supplementary Fig. 8b and Supplementary Table 15 for detailed breakdown), highlighting areas that need further refinement. Finally, 81% of the citations provided by the IR agent were found to be relevant and verifiable. These evaluations by human experts provide evidence that including predicted IR status enhances the quality, trustworthiness and clinical utility of LLM-generated responses to queries about metabolic health, forming a solid foundation for the future development and deployment of similar AI-driven health assistants.
Discussion
Using a cohort of 1,165 participants, our proposed IR prediction framework represents the first, to our knowledge, deployable end-to-end model that uses readily available data from wearables, demographics and routine blood biomarkers. The models were trained using a ground-truth measure of IR (HOMA-IR) that has been validated and established in previous large epidemiological studies3. In the USA, 26% of the population own smartwatches, and approximately 15% undergo annual medical exams that include routine blood biomarker assessments. These statistics are expected to increase with wider adoption of wearables and growing recognition of the health benefits associated with continuous health monitoring. Although many studies have developed predictive models for IR using demographic and clinical biomarkers29,30,31,32, integrating data from wearable devices has not been considered. Individual lifestyle factors, such as physical activity, influence IR considerably, making the potential of wearable devices to enhance IR prediction a crucial area of study.
A key contribution of this work is the use and fine-tuning of a WFM that learns robust, high-dimensional representations directly from high-resolution sensor data. Our results show that this approach significantly outperforms conventional methods that rely on simple aggregate metrics. The WFM not only improved the predictive accuracy across all multimodal combinations, but it also substantially increased the feature importance of the wearable-device data, indicating that it successfully captures the complex physiological dynamics of IR that are missed by simpler approaches. Crucially, this advantage was not limited to cross-validation; the WFM-enhanced models showed strong generalizability when tested on an entirely independent validation cohort, maintaining their superior performance on unseen data. Therefore, the WFM represents a powerful and generalizable method for unlocking the full predictive potential of data from wearable sensors in assessing metabolic health.
The gold-standard test for IR, the hyperinsulinaemic euglycaemic clamp2, is impractical for routine clinical use because of its complexity and resource intensiveness. At present, HOMA-IR is not routinely assessed, owing to the cost and logistical challenges of insulin testing. Whereas a single insulin measurement can be performed at a reasonable cost in clinical settings, routine insulin assessment presents logistical (clinical laboratory visits) and cost-related challenges. At-home testing is unlikely to mitigate these logistical hurdles, because standard insulin immunoassays are not yet readily adaptable to user-friendly, home-based microfluidic kits. Consequently, repeated insulin testing necessitates ongoing visits to a clinical laboratory, with substantial cumulative costs. Therefore, our model using readily available digital data and blood biomarkers could serve as a screening tool to prioritize individuals whose insulin levels should be tested in a clinical laboratory setting to calculate their exact HOMA-IR.
Previous research involving wearable smartwatches and IR focused mainly on investigating associations between wearable-derived features (for example, RHR and HRV) and IR, without developing predictive models20,21,22,23. The most recent state-of-the-art method for detecting IR at home involves performing two OGTTs at home using CGM to analyse glucose time-series data. These methods have yielded an AUROC of 0.88 (ref. 18). Although the ability to infer IR is a crucial step towards developing widely accessible diagnostic tests, the adoption of CGMs among people who are not diabetic remains limited. Therefore, our approach, which uses wearable smartwatch devices, demographic data and readily available blood markers, offers a more scalable solution that does not require additional testing. With regard to blood biomarkers, although numerous studies have found strong correlations between HOMA-IR and biomarkers such as triglycerides and HDL33,34,35,36, few have aimed to predict IR using readily available blood markers. Existing models have shown suboptimal predictive performance, relied on small sample sizes or included insulin itself as a predictor, a marker not typically included in routine annual exams37.
Previous studies used anthropometric measures (for example, BMI, waist circumference and so on) or lifestyle surveys (for example, diet and physical activity) to predict IR. However, these methods generally performed suboptimally, mainly because they failed to capture underlying physiology or to provide continuous monitoring of lifestyle factors32,38. Notably, one group proposed logistic regression models for IR prediction, using anthropometric measures and blood biomarkers29. Using a paediatric cohort from Portugal, and defining IR with a HOMA-IR threshold of 3.4 (the 95th percentile in healthy Mediterranean children and adolescents), their model—which incorporated BMI, obesity duration, brachial, waist and hip circumferences, acanthosis nigricans, Tanner stage, self-reported physical activity, family history of T2D, hypertension and fasting glucose—achieved a sensitivity and specificity of 0.816 in the initial cohort. However, in the validation cohort, the sensitivity was 0.81 and the specificity was 0.52. By contrast, our model—which uses demographics (only BMI and age), physical activity measured passively and objectively through wearable-device features, and fasting glucose—achieved a specificity of 0.84 and a sensitivity of 0.73 in the initial cohort, and a specificity of 0.83 and a sensitivity of 0.79 in the independent validation cohort. Waist circumference emerges as the top anthropic metric that is associated with IR. Waist index, calculated as waist circumference (cm) divided by 94 for men and 80 for women, has shown a strong correlation with HOMA-IR39, and it was used to predict IR (HOMA-IR > 2.5), with a sensitivity of 0.78 and specificity of 0.65. In our study, we first used a HOMA-IR threshold of 2.9, which yielded a sensitivity of 0.78 and specificity of 0.84. However, after recalculating sensitivity and specificity using a threshold of 2.5, we obtained a sensitivity of 0.82 and specificity of 0.78. Our method provides a scalable initial screening tool to identify individuals who have an increased likelihood of IR. Positive screening results would prompt referral for clinical fasting glucose and insulin assays, allowing accurate HOMA-IR calculation and subsequent clinical discussion. Consequently, a model with higher specificity than that of existing surrogates is essential to minimize unnecessary testing in laboratory settings.
Several studies have associated lifestyle intervention programmes, such as weight loss and increased aerobic exercise, with significant reductions in IR40,41. Future longitudinal studies should examine whether weight loss in individuals leads to decreases in IR that are detectable by wearable devices. In addition to longitudinal validation, simulations could model the magnitude of lifestyle changes needed to improve IR and the degree of change required to reverse the progression of diabetes42,43.
Although our study included diverse participants in terms of age and gender, only 25% of participants had complete data and were included in the analysis. The final cohort might have overrepresented those with cardiometabolic disease or health awareness owing to the requirement for laboratory blood tests and use of wearable devices. Moreover, genetics and the microbiome are established contributors to IR44,45. Future studies would benefit from collecting and analysing these data in relation to IR prediction models. This will help to elucidate the interplay between lifestyle and genetic factors in IR.
We recognize that the current adoption of advanced wearables is skewed towards Caucasian individuals. However, basic wearable fitness trackers with the necessary measurement capabilities are becoming increasingly accessible, and we expect this trend to continue. In our models, we incorporated features that are commonly found on lower-cost devices, ensuring that our models are applicable to a wide range of devices, and maintaining their relevance and practical utility. In our study, we used data from seven smartwatches and four trackers, all from the same manufacturer (Google, which owns Fitbit) and all using similar algorithms. Although we expect the results to generalize to other brands, further investigation and specific quality-control steps to mitigate known accuracy issues are warranted.
Our training of the machine-learning models relied on HOMA-IR as a proxy to scale the experiments to thousands of participants, instead of relying on the gold-standard hyperinsulinaemic euglycaemic clamp, which is laborious and costly. However, using HOMA-IR as a ground truth presents some challenges, mainly related to per-subject reproducibility and standardization of insulin concentrations across laboratories. This can lead to a reported coefficient of variation of 23.5% between two measurements46. In our study, all participants underwent blood tests at Quest Diagnostics branches across the USA. This approach minimizes the risk of variation in insulin measurement standardization across different laboratories, and hence reduces HOMA-IR variability. Although we acknowledge this limitation, we emphasize that HOMA-IR remains a valuable tool for classifying individuals into broad categories of insulin sensitivity or resistance, rather than for providing precise quantifications of resistance levels. Therefore, our framework prioritizes the evaluation of IR class over the prediction of specific HOMA-IR values.
In the future, IR detection could be used as a more objective measure for prescribing GLP1 and GIP receptor agonists. At present, providers rely on guidelines based on BMI to determine the need for these medications47, but BMI is not an ideal metric to use to differentiate between people who require such drugs through medical necessity and those who choose to take them for lifestyle purposes. An IR measure would provide both providers and payers with a more robust and objective assessment when prescribing and reimbursing for these drugs. However, our proposed IR detection models need further fine-tuning to improve their sensitivity and specificity, particularly in the context of identifying subpopulations who would benefit most from incretin-hormone-based therapies.
Methods
Remote recruitment of participants through the GHS platform
The study protocol was approved by Advarra (IRB no. Pro00074093). Participants were recruited using the Google Health Services (GHS) and the Fitbit applications. GHS is a platform for running digital studies that allows participants to enrol, check eligibility and provide informed consent. GHS enables the collection of data from wearables (Fitbit devices or Pixel watches) and allows participants to complete questionnaires and order blood tests through Quest Diagnostics. In addition to the consent, participants were asked to sign the Quest HIPAA authorization as part of the consent flow within the GHS app. This study resulted in the enrolment of 4,416 participants in the USA, a subset of whom (n = 1,165) had complete data and were therefore included in our analysis. The study was performed remotely with one visit to a Quest Patient Service Center for a blood draw. Participants were asked to wear their wrist-worn devices continuously, schedule and complete a blood draw and answer questionnaires.
Inclusion criteria included: participants residing in the USA aged between 21 and 80; Android Fitbit users who wear a Fitbit device or a Pixel watch with heart-rate-sensing capabilities; users who have at least three months of existing data in which they have used the device for at least 75% of the days to track their activity and sleep; participants willing to update their Fitbit Android app; participants willing to install or update their GHS app on their Android phone; participants willing to link their Quest Diagnostics account to the GHS app (or create an account); participants able to speak and read English and provide informed consent and HIPAA authorization to take part in the study; and participants with access to and who are willing to go to a Quest Diagnostics location for blood draws.
Exclusion criteria included participants living in Alaska, Arizona, Hawaii and the US territories, because Quest Diagnostics cannot provide blood tests in those states; participants with uncontrolled disease (for example, a recent change in treatment in the past six weeks, awaiting review to trigger a change of treatment, a treating physician has indicated the condition is not yet controlled, or where symptoms of the condition are not responding to treatment); and participants with conditions that might make collecting blood samples through venipuncture impractical.
Study design
As part of this study, participants were asked to link their Fitbit account with the GHS app. They were also asked to grant GHS permission to collect Fitbit data throughout the study, including data for up to three months before study enrolment. Once participants were enrolled in the study they were asked to do the following: (1) wear their Fitbit device or Pixel watch during the day and while they sleep (at least three out of every four days) for the duration of the study; (2) complete four questionnaires, which covered (i) demographic information, (ii) health history and health information, such as sleep and exercise habits, (iii) participant perception of health, and (iv) blood test interpretation (see below for details); (3) schedule an appointment to complete the laboratory test orders that were placed for them and go to a Quest Patient Service Center for a blood draw within 65 days of enrolment; (4) complete a blood draw at the Quest Patient Service Center; (5) review blood test results in the GHS app when available.
Collection of metadata
Demographics (for example, age, gender, ethnicity, weight and height), optional measures, such as medical conditions (for example, diabetes, hyperlipidaemia, cardiovascular disease, kidney disease, hypertension and so on), blood pressure, waist circumference, medications, self-reported health management and habits were collected through a baseline questionnaire that participants completed immediately after enrolment through the GHS app.
Blood biomarker measurements at Quest Diagnostics
Eligible participants were asked to schedule and complete a visit to a Quest Diagnostics Patient Service Center of their choice in their local area. This visit included a standard blood draw to measure the following: complete blood count, CMP, insulin, total cholesterol, triglycerides, HDL cholesterol, calculated LDL cholesterol, HbA1c, high-sensitivity CRP, hepatic panel, gamma-glutamyl transferase (GGT) and total testosterone. For this research study, we only had access to this predefined set of laboratory tests and did not collect or receive any blood test data not included in this study. Participants were asked to have their blood drawn while fasting for at least eight hours, in the early morning, 07:00–10:00 local time, to minimize the effect of the solar diurnal cycle. This study also had clinical oversight by a physician network partner. Laboratory results from the blood draw were returned to participants and made available for participant review in the GHS app for the duration of the study; however, these were pulled directly from Quest Diagnostics each time this was requested by the participant and were not stored in the GHS app. Participants provided consent and HIPAA authorization to grant GHS permission to collect the corresponding results from Quest Diagnostics. Data transferred from Quest Diagnostics were retrieved securely using encrypted protocols. Once the study was completed, laboratory results remained in the participant’s Quest Account in accordance with Quest’s standard practices.
Data from wearables (Fitbits and Pixel watches)
Participants were asked to wear their own Fitbit or Pixel watch, which can include sensors and metrics such as photoplethysmogram (PPG), gyroscope, altimeter, accelerometer, on-wrist skin temperature, blood oxygen saturation (SpO2) and ambient light sensor. Participants consented to share the following data from these devices: (1) heart-rate metrics: heart rate, RHR captured daily, interbeat interval (IBI, also known as RR interval) calculated from the PPG sensor, and HRV metrics (such as RMSSD, standard of normal-to-normal intervals, standard deviation of RR intervals, percentage of successive normal-to-normal heartbeats that differ by more than 50 ms, and so on); (2) physical activity metrics: steps, floors and active zone minutes (AZMs); (3) sleep metrics: bed time, wake time, sleep duration, sleep stages, sleep quality, sleep coefficients and sleep logs; (4) respiration and skin temperature: respiration rate values during the day and night and skin temperature (if the sensor is available on the wearable); (5) blood oxygen saturation (SpO2) values during the day and night (if available on the wearable); (6) weight: measure of weight that may be logged in the Fitbit account; and (7) exercise and activity data: daily total exercise sessions completed and logs of activities that have been classified. Fitbit daily RHR is calculated from periods of stillness throughout the day, as determined by the on-device accelerometer. If a person wears their device while sleeping, their sleeping heart rate is also included in the calculation. Daily RMSSD HRV is calculated from pulse intervals measured during sleep periods greater than three hours. AZMs is a feature that tracks the time a person spends in different heart-rate zones during physical activity. A person receives one AZM for every minute spent in the ‘moderate’ zone, and two AZMs for every minute spent in the ‘vigorous’ or ‘peak’ zones. The heart-rate zones are based on percentage of heart-rate reserve achieved, in which the heart-rate reserve is the difference between the maximum heart rate and the RHR. The moderate zone is defined as 40–59% of heart-rate reserve, vigorous as 60–84% and peak as 85–100%.
Selection of HOMA-IR thresholds for IR
Our selection of HOMA-IR thresholds was based on a previous study24, which reported a HOMA-IR range of 1.5 to 3 for IR. We chose a HOMA-IR threshold of 2.9, approximating the midpoint between the NHANES-derived threshold for the US population (HOMA-IR = 2.77) and the maximum value in the review (HOMA-IR = 3). For insulin sensitivity, we used the lowest threshold from the review, HOMA-IR = 1.55, which was rounded down to 1.5. Participants with HOMA-IR values between 1.5 and 2.9 were classified as impaired-IS.
Modelling and computational pipeline
Our method consists of four stages: (i) data preprocessing; (ii) modelling and training; (iii) prediction and classification of HOMA-IR; and (iv) LLM-based interpretation of the results. We describe each of these components, including evaluation strategies and large-scale ablation studies, below.
Data preprocessing
Age and BMI (demographics)
From the user-provided data on recruiting surveys, we extract a user’s age and compute BMI from height and weight. As a quality-control process, we exclude users with a BMI greater than 65, or lower than 12.
Digital markers derived from wearables (wearable features)
Using the estimated digital markers from Fitbit algorithms, we aggregated users’ digital markers using mean, standard deviation and median values for the past n = {7, 14, 30, 60, 60} days before blood test collection. We performed ablation studies to find the ‘optimal’ value of n (described later in this section).
Biomarkers from blood biochemistry (blood tests)
As a first filter, we removed any participant who was not fasting as reported in a survey at the time of blood test collection. In addition, for each experiment, we excluded participants with missing values from the input feature set. Lastly, to remove outliers, we used the true HOMA-IR value (HOMA-IR = (fasting insulin (µU ml−1) × fasting glucose (mg dl−1))/405) and excluded any participant whose HOMA-IR value was greater than or equal to 15.
Data standardization
The data used for modelling are a concatenation of demographics, aggregated digital markers and blood biomarkers. To create consistent modelling data that were agnostic to the learning model, we standardized input features to have zero mean and unit variance. For each training fold, our ‘normalizer’ object was fitted to the data in the training subset (not including samples in the testing subset). The fitted object was then used to transform the samples in both the training and the testing subset. The standardized data were used for all modelling tasks and evaluations.
To test the generalizability of our approach to the validation cohort, we used the same normalizer object that was fitted to the training data from the initial cohort.
Modelling
To determine the risk of IR, our goal was first to predict the value of HOMA-IR, and use existing thresholds for classifying individuals. Performing regression before classification allows for flexibility and much greater interpretation of our results and analysis of individual data points.
Direct regression
As our first approach in modelling HOMA-IR, we used gradient boosting machines; specifically, the XGBoost framework48,49. Gradient boosting methods excel in handling complex datasets with potentially non-linear relationships, making them particularly well-suited for our task. Compared with conventional tree-based methods, XGBoost provides computational efficiency, scalability and regularization techniques that enhance model generalization. Although it is often associated with tree-based models, XGBoost offers a versatile framework for making use of weak learners, which include both linear and tree-based learners. Owing to the ambiguity that surrounds the linear interaction between various input features, we used both linear (L1–L2) and non-linear (tree-based) learners to assess the complexity of the problem space.
Given n data-label pairs (xi, yi), gradient boosting can be viewed as an additive combination of K weak learners f, with the aim to predict target \({\hat{y}}_{i}\) through:
$${\hat{y}}_{i}=\mathop{\sum }\limits_{k}^{K}{f}_{k}({x}_{i}),{f}_{k}\in {\mathcal{F}}.$$
(1)
\({\mathcal{F}}\) denotes the space of all possible regression trees, \({\mathcal{F}}=\{f(x)={w}_{q}(x)\}\), with \(w\in {{\mathbb{R}}}^{T}\), where \(T\) is the number of leaves in the tree, and \(q:{{\mathbb{R}}}^{d}\to T\) represents the structure of each tree that maps d-dimensional data points (that is, \({x}_{i}\in {{\mathbb{R}}}^{d}\)) to the corresponding leaf index. In our notation, fk(xi) represents the prediction made by the k-th tree in the ensemble for the input sample xi. Although the choice of learners f can determine the linearity of the model, the number of trees and number of leaves serve as key hyperparameters that can further control model complexity. The objective of extreme gradient boosting machines is to minimize the loss function
$${\mathcal{L}}(\phi )=\mathop{\sum }\limits_{i}^{n}l({\hat{y}}_{i},{y}_{i})+\mathop{\sum }\limits_{k}^{K}\varOmega ({f}_{k}),$$
(2)
where \(l(\,{\hat{y}}_{i},{y}_{i})\) represents the (traditional gradient boosting machines) loss function measuring the discrepancy between the predicted output \(\hat{{y}_{i}}\) and yi, and the regularizer Ω(fk) penalizes model complexity, where \(\varOmega =\gamma T+\frac{1}{2}\lambda \Vert w{\Vert }^{2}\), with \(\gamma \) and \(\lambda \) being hyperparameters.
In our work, the first set of models use weak linear learners that can capture additive linear relationships within the data. This approach assumes that the target variable can be modelled as a linear combination of the input features, potentially revealing important feature interactions (this is obtained by using gblinear from the XGBoost implementation50). Recognizing the potential for complex non-linear relationships within our data, we let the second set of models be non-linear (through gbtree in the XGBoost implementation). XGBoost’s efficient tree construction algorithm, based on a pre-sorted split finding algorithm and sparsity-aware split finding, allows for faster training and exploration of a larger parameter space. This, coupled with the inherent ability of decision trees to model non-linear interactions, make the tree-based approach a powerful technique for uncovering potential complex patterns in our study.
Representation learning
Although direct regression methods, such as XGBoost, can effectively handle high-dimensional data, we hypothesized that their performance can be further amplified by providing informative and concise data representations. Studies have demonstrated the importance of mathematical representation of personal health records (blood tests, lifestyle data and so on) for various downstream tasks51. To test the representation learning hypothesis, we used representation learning techniques (specifically, MAEs), potentially learning latent representations that could enhance regression performance.
MAE
Although simple autoencoders (AEs) have been shown to be an effective unsupervised approach for representation learning, MAEs52, a self-supervised variant of AEs, have shown tremendous improvements over conventional AEs, achieving state-of-the-art results across different benchmark datasets52,53. For our approach, given a datapoint \({x}_{i}\in {{\mathbb{R}}}^{d}\), we start by drawing a masking vector \(m\in {{\mathbb{R}}}^{d}\) from a multivariate Bernoulli distribution with probability p (determined through ablation studies, as discussed later). Using the mask vector, we generate a masked version of the input, \(\hat{{x}_{i}}=x\odot m\), which is then inputted to the MAE model. Similar to the conventional AE, the objective of our network is to reconstruct the initial input, but from the masked vector \({\hat{x}}_{i}\) as opposed to xi. Therefore, our object for the MAE can be expressed as:
$${{\mathcal{L}}}_{{\rm{AE}}}(x)={{\mathbb{E}}}_{x}\parallel {\rm{Dec}}({\rm{Enc}}(x\odot m))-x{\parallel }^{2}+\lambda \cdot {\rm{SL}}(x,{\rm{Dec}}({\rm{Enc}}(x\odot m))).$$
(3)
Model training and HOMA-IR prediction from learned representations
The results presented for regression in this study include two stages: (i) representation learning and (ii) using the learned representation to train a XGBoost with linear learners for predicting HOMA-IR values. We chose using linear XGBoost models for the second stage to better showcase the improvements made possible by our representation learning approach.
We trained both models for a total of 500 epochs with the smooth L1 loss coefficient λ = 0.01. For optimization, we used the Adam optimizer with (β1, β2) = (0.9, 0.999) and ϵ = 10−12. The initial learning rate was set to lr = 0.001, scheduled to decrease exponentially (γ = 0.95) after 100 epochs on a plateau with a patience of 20 epochs. For the MAE model, we set the masking probability p = 0.75 based on our empirical results (found through grid search) and existing literature.
Parameter selection
To identify the set of parameters used for the ablation studies, we performed grid search for key parameters. For all models, because of grid search’s exponential computational complexity, we performed grid search on a common feature set and time window with fivefold cross-validation to identify optimal hyperparameter values. We describe the parameter grid for each model in Supplementary Table 16. To make our experiments deterministic, our random seed was set to [0, 92, 1, 2024, 12121].
WFMs
Recent years have seen the rise of ‘foundation models’, whereby large-scale pretraining on diverse—and often unlabelled—data has led to representations that are useful for a breadth of downstream applications27,28. To evaluate the utility of such large-scale pretraining on the task of IR prediction, we used the large sensor model (LSM)54,55, a foundation whose embeddings have shown utility for a number of health downstreams, including activity recognition, mood classification, hypertension detection and anxiety detection. Specifically, for the IR prediction, we pretrain an LSM-2-style WFM on 40 million person-hours sampled at minute data resolution from Fitbit users. Each data sample contains 26 minutely aggregated features from a set of 5 sensors (photoplethysmography, accelerometer, skin conductance, altimeter and temperature) for a time span of 1,440 min (one day). The 26 features used as input for model embedding function features are described in Supplementary Table 17.
A core property of these data is that they have complex, structured missingness patterns. Missing data are ubiquitous in long-duration wearable sensor recordings, with 0% of samples over our entire dataset of 1.6 million instances of one-day data having 0% missingness. Missingness is also complex, originating from a variety of causes with varied structures. For example, large chunks of the signal can be missing if a sensor is temporarily off to save battery life; full missingness across all channels can occur in periods in which the device is not worn (off body); and short bursts of missingness can exist as a result of filtering out measurements that are clearly spurious or out of range.
In the pretraining objective, non-masked tokens are encoded before masked tokens are reintroduced for reconstruction in the subsequent decoder layers. Inputs are tokenized with a one-dimensional (1D) patch size of 10 min, resulting in 3,744 tokens (144 tokens per signal). This is implemented with a shared kernel across channels, using a two-dimensional (2D) positional embedding to encode time and signal identity. Then, the tokens are passed into ViT-1D with 25 million parameters, 12 encoder layers, with a 384-dimensional hidden size, and 4 decoder layers, with a 256-dimensional hidden size.
The pretraining procedure is performed on 8 × 16 Google v5e TPUs with a batch size of 512 for 100,000 steps. During pretraining, we include the inherited mask as well as a mix of artificial masking strategies. The inherited mask is on top of the pre-existing missingness inherent to the data, and the artificial mask is introduced onto present data to have ground truths for reconstruction loss. Our artificially masking mix seeks to model the real-world missingness patterns. The mix includes (1) 80% random imputation masking (to model noise), in which a random patch is masked; (2) 50% temporal slice masking (to model off body), in which all sensors at a random time point are masked; and (3) 50% signal slice masking (to model sensor off), in which all time points for a random sensor are masked. Each instance uses a randomly selected masking strategy with equal probability. Notably, we do not back-propagate on inherited mask tokens.
During evaluation, the pretrained model is then able to operate directly on incomplete multimodal sensor data by dynamically attending only to observed segments. This eliminates the need for external preprocessing, such as imputing or discarding missing values, and ensures generalization from pretraining to downstream deployment in real-world settings. In this way, to obtain our frozen embedding for the XGBoost head to be applied, we take the output of the missingness-agnostic encoded representation and take an average pooling across all unmasked tokens.
Independent validation cohort
The study protocol was approved by WCG (IRB no. 1371945). This validation cohort is part of a larger, 30-week longitudinal study designed to assess the effects of lifestyle modifications on cardiometabolic health metrics. Data collected throughout the study included anthropometric measurements (for example, BMI, waist circumference, skin tone and blood pressure), blood biomarkers (such as HbA1c, fasting glucose, fasting insulin and lipid panel), wearable-device data (Fitbit Charge 6, measuring heart rate, RHR, HRV, sleep duration and step count) and health and lifestyle questionnaires. Participants attended in-person appointments at the Fitbit Human Research Laboratory in San Francisco at the beginning and end of the study to complete the aforementioned measurements, tests and surveys. All participants were required to wear a Fitbit Charge 6 device continuously for a minimum of 20 h per day throughout the study.
The inclusion criteria of this study included: participants must be located in California, be at least 18 years of age, be able to stand and walk without aid, be willing to provide informed consent, be comfortable with English instructions, own and use a smartphone, be willing to download and use the Fitbit app, be comfortable using wearable devices, have regular internet access and be willing to comply with study procedures. Exclusion criteria included: unwillingness to attend laboratory visits; having uncontrolled hypertension or recent changes in related medications; conditions making blood draws impractical; known bloodborne pathogens; T1D; T2D with HbA1c > 8.0%; history of gastric surgery; non-iron-deficiency anaemia; severe mental illness; heart failure; undergoing dialysis; terminal illness; taking medications for diabetes or glucose control (with specific examples provided); taking oral steroids; using tanning lotions on arms, having wrist tattoos; skin conditions that interfere with devices; pregnancy or plans for pregnancy; recent radiological procedures with contrast agents; having non-permissible internal or irremovable external objects; weight greater than 450 lbs (204 kg); and unwillingness to participate in test protocols.
To start, 144 individuals were enrolled; 127 individuals had complete wearable-device data and 82 individuals had complete blood biomarker data acquired during an in-person visit at the end of the study. Ultimately, 72 individuals had both complete wearable-device data and complete physiological biomarker data, and this group was used as an independent validation cohort for the IR prediction model. For the validation, we used blood biomarkers from the final study time point to ensure the corresponding wearable-device data constituted a substantial representation of participants’ lifestyles. Wearable-device data encompassed all available records up to the final visit. Similarly, demographic data (age and BMI) were extracted from the final visit.
Evaluation of prediction and classification of HOMA-IR
Evaluation of regression
The first stage of our framework for predicting IR is to predict the continuous HOMA-IR values (regression). To evaluate the performance of each experiment, we stored the predicted HOMA-IR values in the test set of each fold, computing the average and standard deviation of mean absolute error across all test folds. To compute the over-concordance between our predicted values and the true HOMA-IR, we calculated the R2 on all predicted values, which includes the prediction for all individuals for when they were included in the test set of each fold.
Evaluation of IR classification
To classify an individual as insulin resistant, we use continuous HOMA-IR values (either predicted or true values for ground truth) and use the threshold of IR = 2.9 for binary classification. Individuals with a HOMA-IR value greater than or equal to 2.9 are considered to be insulin resistant, and are not insulin resistant otherwise. To evaluate our classification, we used the metrics specificity, sensitivity, precision, AUROC and area under precision-recall curve (AUPRC). Similar to the regression evaluation, we report the average and standard deviation of these metrics across each test fold for each experiment.
Time-dependent sensitivity and robustness analysis
To determine the robustness of our approach to time-dependant aggregation, we performed n-day aggregation of time series on a rolling window for the entire study duration. The goal of this analysis was to check the consistency (robustness) of our predictions for various time windows within the study. We report the consistency of predictions for all individuals for n = {7, 14, 30, 60}. We did not include 90 and 120 days, because these windows, maximally, would result in one or two windows over the study period. In addition, we compute the coefficient of variation of the predicted HOMA-IR values across the windows and report the results.
IR agent
LLMs are powerful tools for generating language, and have considerable potential to improve human–computer interactions across many domains, including education, clinical practice and interpretation of personal health metrics56,57. Given the known challenges for patients of understanding laboratory results and derived metrics, such as HOMA-IR, we aimed to develop an LLM-based framework for interpreting the results, as well as interacting with users for follow-up questions and recommendations.
Our proposed LLM-based agent is a ReAct (reasoning and acting) agent58 that is capable of multi-step reasoning and planning. ReAct agents are a class of LLM-based agents that synergistically combine internal reasoning with external actions to accomplish complex tasks. Unlike traditional LLMs, which mainly generate text, ReAct agents interleave textual reasoning traces (‘thoughts’) with calls to external tools or information sources (‘actions’) and reasoning over this multi-step process (‘observe’).
Our IR agent uses the ReAct iterative process to dynamically plan, gather necessary information (for example, from the web), adjust its strategy on the basis of retrieved facts or computed observations and ultimately respond to the incoming health-related query. The IR agent’s ‘thoughts’ provide interpretability and allow the agent to decompose complex problems, whereas its ‘actions’ provide grounding and interaction with the real world or specialized knowledge bases, overcoming the limitations of relying solely on the pretrained knowledge of the LLM. This interplay between internal deliberation and external interaction enables our agent to tackle tasks that require both reasoning and grounded knowledge acquisition in real time.
IR agent toolbox
A key part of our IR agent is its ability to intelligently select the set of tools it requires for answering incoming queries. The IR agent’s toolbox includes grounding tools (organic web search via Google Search), arithmetic tools, Python sandbox to execute code, as well as our HOMA-IR prediction models. These models and functions serve as a set of actions for our agent, ensuring accurate and deterministic computations that reduce the risk of error and hallucinations58. We describe the set of tools in Supplementary Table 18.
Implementation of the IR agent
We use Google DeepMind’s open-source OneTwo library59 to implement the IR agent. OneTwo is a Python framework specifically designed for facilitating research on prompting strategies for large foundation models. Crucially, OneTwo’s flexible execution model supports the complex interaction patterns required by our IR agent, enabling the interleaving of LLM-generated reasoning (‘thoughts’) with external ‘actions’ such as web search or executing our trained machine-learning models. OneTwo’s built-in support for tool use and agent abstractions directly facilitated the development of an agent that is LLM-agnostic (that is, it can use other LLMs, not just models developed by Google).
Evaluation of the IR agent
To evaluate the IR agent, our goal was to measure the added benefit of including information about IR (as predicted by our machine-learning models) for metabolically relevant queries, when appropriate, compared with LLM responses that do not include this information. In addition, we wanted to assess the accuracy of the IR interpretation and comprehensiveness in these responses, as evaluated by endocrinologists (human experts).
Set-up of the human expert evaluation
From our study, we selected five participants with atypical metabolic profiles, in consultation with a doctor who was not part of the evaluation team, to ask metabolically relevant questions.
These participants were selected from the following groups:
-
Individuals with clinically normal HbA1c, but classified as insulin resistant;
-
Individuals with obesity with a sedentary lifestyle (median steps of fewer than 8,000 and average steps of fewer than 8,000 steps per week) who were classified as being insulin resistant;
-
Individuals with obesity with an active lifestyle (average step count of 10,000 or more per day, computed on a weekly basis) who were classified as insulin resistant;
-
Individuals with obesity with an active lifestyle who were classified as insulin sensitive.
Using the data from these individuals, we then generated responses using our IR agent and the same base LLM at the core of our agent (Gemini 2.0 Flash); note that both models had access to the same data, which include demographics, blood biomarkers and wearable features, and were asked the same questions.
After generating answers to these questions from the IR agent and the base LLM, we recruited five practicing endocrinologists to evaluate each model response in a blind manner, in which the expert evaluators did not know which response was from the base model and which from the proposed IR agent, and the design or the scope of the IR agent was not shared with them to avoid any potential bias in the the ratings. More specifically, we asked the endocrinologists to evaluate the response in two manners:
Side-by-side comparison
This technique is commonly used in the field to measure the effect of variables (for example, inclusion or exclusion of certain features) in generated responses from similar models. The aim was to measure the benefits provided by our IR agent, including the addition of IR information, compared with the base LLM without access to the IR information and the specialized tool. We asked the expert evaluators to answer the following rubrics while considering responses side-by-side:
-
Q1 [Comprehensiveness]. Which of the two responses are more complete or more comprehensive? (In your opinion, which is a better response?)
-
Q2 [Trustworthiness]. Which response do you find more trustworthy?
-
Q3 [Personalization]. Which response better personalizes different aspects of health (for example, lifestyle and cardiovascular health) with respect to the query?
Absolute accuracy of the IR agent
In addition to the side-by-side comparisons, we aimed to evaluate the accuracy of the IR agent by itself across different clinically relevant dimensions60. Therefore, we asked the endocrinologists to rate the following ‘absolute’ rubric questions:
-
Q1 [Factuality]. Are all the general statements in the response (not related to a user’s specific data or condition) factually accurate?
-
Q2 [Reference and Interpretation]. Does the response reference the user’s personal data and interpret it correctly?
-
Q3 [Safety]. Is the response free from potentially harmful medical advice or recommendations that if acted upon may cause harm to the user?
-
Q4 [Grounding]. If provided citations, are all citations in the response from relevant and verifiable sources?
Statistical analysis
Statistical significance was determined using two-sided Wilcoxon rank-sum tests, with P values adjusted for multiple comparisons using the Benjamini–Hochberg method. Data processing, model training and evaluation were implemented in Python using numpy v.2.0.2, tensorflow v.2.19.0, scipy v.1.16.3, statsmodels v.0.14.6, sklearn v.1.6.1, shap v.0.50.0, xgboost v.3.1.2, torch v.2.9.0, pandas v.2.2.2, umap v.0.5.9.post2, pickle v.4.0, pytz v.2025.2, re v.2.2.1, tqdm v.4.67.1, IPython v.7.34.0, json v.2.0.9 and altair v.5.5.0.
Visualization methods
We used matplotlib v.3.10.0, seaborn v.0.13.2, bokeh v.3.7.3 and Figma to plot most of the figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The de-identified dataset used in this study is available to approved researchers for reproducibility purposes only. To become an approved researcher, please follow the instructions at https://github.com/Google-Health/consumer-health-research/tree/main/insulin_resistance_prediction.
Code availability
References
Parcha, V. et al. Insulin resistance and cardiometabolic risk profile among nondiabetic American young adults: insights from NHANES. J. Clin. Endocrinol. Metab. 107, e25–e37 (2021).
DeFronzo, R. A., Tobin, J. D. & Andres, R. Glucose clamp technique: a method for quantifying insulin secretion and resistance. Am. J. Physiol. 237, E214–E223 (1979).
Matthews, D. R. et al. Homeostasis model assessment: insulin resistance and β-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28, 412–419 (1985).
International Diabetes Federation. Facts & figures. https://idf.org/about-diabetes/diabetes-facts-figures/ (2022).
Galaviz, K. I., Venkat Narayan, K. M., Lobelo, F. & Weber, M. B. Lifestyle and the prevention of type 2 diabetes: a status report. Am. J. Lifestyle Med. 12, 4–20 (2018).
El, K. et al. GIP mediates the incretin effect and glucose tolerance by dual actions on α cells and β cells. Sci. Adv. 7, eabf1948 (2021).
Roep, B. O., Thomaidou, S., van Tienhoven, R. & Zaldumbide, A. Type 1 diabetes mellitus as a disease of the β-cell (do not blame the immune system?). Nat. Rev. Endocrinol. 17, 150–161 (2020).
Sapra, A. & Bhandari, P. StatPearls: Diabetes (StatPearls Publishing, 2022).
Bonora, E. et al. Prevalence of insulin resistance in metabolic disorders: the Bruneck Study. Diabetes 47, 1643–1649 (1998).
Turcotte, L. P. & Fisher, J. S. Skeletal muscle insulin resistance: roles of fatty acid metabolism and exercise. Phys. Ther. 88, 1279–1296 (2008).
Niemann, M. J., Tucker, L. A., Bailey, B. W. & Davidson, L. E. Strength training and insulin resistance: the mediating role of body composition. J. Diabetes Res. 2020, 7694825 (2020).
Zhang, X. et al. Impacts of selected dietary nutrient intakes on skeletal muscle insulin sensitivity and applications to early prevention of type 2 diabetes. Adv. Nutr. 12, 1305–1316 (2021).
Demaria, T. M. et al. Once a week consumption of Western diet over twelve weeks promotes sustained insulin resistance and non-alcoholic fat liver disease in C57BL/6 J mice. Sci Rep. 13, 3058 (2023).
Ko, K. D., Kim, K. K. & Lee, K. R. Does weight gain associated with thiazolidinedione use negatively affect cardiometabolic health? J. Obes. Metab. Syndr. 26, 102–106 (2017).
Sui, Y. et al. Long-term treatment with metformin in the prevention of fatty liver in Zucker diabetic fatty rats. Diabetol. Metab. Syndr. 11, 94 (2019).
Garvey, W. T. et al. Two-year effects of semaglutide in adults with overweight or obesity: the STEP 5 trial. Nat. Med. 28, 2083–2091 (2022).
Frias, J. P. et al. Tirzepatide improved markers of islet cell function and insulin sensitivity in people with T2D (SURPASS-2). J. Clin. Endocrinol. Metab. 109, 1745–1753 (2024).
Metwally, A. A. et al. Prediction of metabolic subphenotypes of type 2 diabetes via continuous glucose monitoring and machine learning. Nat. Biomed. Eng. 9, 1222–1239 (2025).
Hall, H. et al. Glucotypes reveal new patterns of glucose dysregulation. PLoS Biol. 16, e2005143 (2018).
Flanagan, D. E. et al. The autonomic control of heart rate and insulin resistance in young adults. J. Clin. Endocrinol. Metab. 84, 1263–1267 (1999).
Beddhu, S., Nigwekar, S. U., Ma, X. & Greene, T. Associations of resting heart rate with insulin resistance, cardiovascular events and mortality in chronic kidney disease. Nephrol. Dial. Transplant 24, 2482–2488 (2009).
Svensson, M. K. et al. Alterations in heart rate variability during everyday life are linked to insulin resistance. A role of dominating sympathetic over parasympathetic nerve activity? Cardiovasc. Diabetol. 15, 91 (2016).
Saito, I. et al. Heart rate variability, insulin resistance, and insulin sensitivity in Japanese adults: the Toon Health Study. J. Epidemiol. 25, 583–591 (2015).
Gayoso-Diz, P. et al. Insulin resistance (HOMA-IR) cut-off values and the metabolic syndrome in a general adult population: effect of gender and age: EPIRCE cross-sectional study. BMC Endocr. Disord. 13, 47 (2013).
Endukuru, C. K., Gaur, G. S., Yerrabelli, D., Sahoo, J. & Vairappan, B. Cut-off values and clinical utility of surrogate markers for insulin resistance and beta-cell function to identify metabolic syndrome and its components among southern Indian adults. J. Obes. Metab. Syndr. 29, 281–291 (2020).
de Cassia da Silva, C. et al. The threshold value for identifying insulin resistance (HOMA-IR) in an admixed adolescent population: a hyperglycemic clamp validated study. Arch. Endocrinol. Metab. 67, 119–125 (2023).
Bommasani, R. et al. On the opportunities and risks of foundation models. Preprint at https://doi.org/10.48550/arXiv.2108.07258 (2021).
Liang, Y. et al. Foundation models for time series analysis: a tutorial and survey. In KDD ’24: Proc. 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining 6555–6565 (ACM, 2024).
Araújo, D., Morgado, C., Correia-Pinto, J. & Antunes, H. Predicting insulin resistance in a pediatric population with obesity. J. Pediatr. Gastroenterol. Nutr. 77, 779–787 (2023).
Berglund, L. & Lithell, H. Prediction models for insulin resistance. Blood Press. 5, 274–277 (1996).
Park, S., Kim, C. & Wu, X. Development and validation of an insulin resistance predicting model using a machine-learning approach in a population-based cohort in Korea. Diagnostics 12, 212 (2022).
Wedin, W. K., Diaz-Gimenez, L. & Convit, A. J. Prediction of insulin resistance with anthropometric measures: lessons from a large adolescent population. Diabetes Metab. Syndr. Obes. 5, 219–225 (2012).
Simental-Mendía, L. E., Castañeda-Chacón, A., Rodriguez-Morán, M., Aradillas-García, C. & Guerrero-Romero, F. Relationship between elevated triglyceride levels with the increase of HOMA-IR and HOMA-β in healthy children and adolescents with normal weight. Eur. J. Pediatr. 174, 597–605 (2014).
Kim, J.-S., Kang, H.-T., Shim, J.-Y. & Lee, H.-R. The association between the triglyceride to high-density lipoprotein cholesterol ratio with insulin resistance (HOMA-IR) in the general Korean population: based on the National Health and Nutrition Examination Survey in 2007-2009. Diabetes Res. Clin. Pract. 97, 132–138 (2012).
Hirschler, V., Maccallini, G., Sanchez, M., Gonzalez, C. & Molinari, C. Association between triglyceride to HDL-C ratio and insulin resistance in indigenous Argentinean children. Pediatr. Diabetes 16, 606–612 (2015).
Olson, K., Hendricks, B. & Murdock, D. K. The triglyceride to HDL ratio and its relationship to insulin resistance in pre- and postpubertal children: observation from the Wausau SCHOOL Project. Cholesterol 2012, 794252 (2012).
McAuley, K. A. et al. Diagnosing insulin resistance in the general population. Diabetes Care 24, 460–464 (2001).
Lee, S., Bacha, F., Gungor, N. & Arslanian, S. A. Waist circumference is an independent predictor of insulin resistance in black and white youths. J. Pediatr. 148, 188–194 (2006).
Magri, C. J., Fava, S. & Galea, J. Prediction of insulin resistance in type 2 diabetes mellitus using routinely available clinical parameters. Diabetes Metab. Syndr. 10, S96–S101 (2016).
McLaughlin, T. et al. Persistence of improvement in insulin sensitivity following a dietary weight loss programme. Diabetes Obes. Metab. 10, 1186–1194 (2008).
Ryan, B. J. et al. Moderate-intensity exercise and high-intensity interval training affect insulin sensitivity similarly in obese adults. J. Clin. Endocrinol. Metab. 105, e2941–e2959 (2020).
Park, H. et al. High-resolution lifestyle profiling and metabolic subphenotypes of type 2 diabetes. NPJ Digit. Med. 8, 352 (2025).
Wu, Y. et al. Individual variations in glycemic responses to carbohydrates and underlying metabolic physiology. Nat. Med. 31, 2232–2243 (2025).
Zhou, W. et al. Longitudinal multi-omics of host–microbe dynamics in prediabetes. Nature 569, 663–671 (2019).
Lotta, L. A. et al. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat. Genet. 49, 17–26 (2016).
Sarafidis, P. A. et al. Validity and reproducibility of HOMA-IR, 1/HOMA-IR, QUICKI and McAuley’s indices in patients with hypertension and type II diabetes. J. Hum. Hypertens. 21, 709–716 (2007).
European Medicines Agency. Wegovy. EMA https://www.ema.europa.eu/en/medicines/human/EPAR/wegovy (2022).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. In Proc. 31st International Conference on Neural Information Processing Systems 4768–4777 (Curran Associates, 2017).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (ACM, 2016).
XGBoost Python Package v.2.1.1. https://xgboost.readthedocs.io/en/stable/python (DMLC, 2024).
Heydari, A. A., Rezaei, N., Prieto, J. L., Patel, S. N. & Metwally, A. A. Lifestyle-informed personalized blood biomarker prediction via novel representation learning. In 2024 IEEE EMBS International Conference on Biomedical and Health Informatics (BHI) 1–8 (IEEE, 2024).
He, K. et al. Masked autoencoders are scalable vision learners. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 15979–15988 (IEEE, 2022).
Zhang, Q., Wang, Y. & Wang, Y. How mask matters: towards theoretical understandings of masked autoencoders. In NIPS’22: Proc. 36th Conference on Neural Information Processing Systems 27127–27139 (ACM, 2022).
Narayanswamy, G. et al. Scaling wearable foundation models. In Proc. 13th International Conference on Learning Representations 98103–98134 (ICLR, 2025).
Xu, M. A. et al. LSM-2: learning from incomplete wearable sensor data. Preprint at https://doi.org/10.48550/arXiv.2506.05321 (2025).
Saab, K. et al. Capabilities of Gemini models in medicine. Preprint at https://doi.org/10.48550/arXiv.2404.18416 (2024).
Heydari, A. A. et al. The anatomy of a personal health agent. Preprint at https://doi.org/10.48550/arXiv.2508.20148 (2025).
Yao, S. et al. ReAct: Synergizing reasoning and acting in language models. In Proc. 11th International Conference on Learning Representations https://openreview.net/pdf?id=WE_vluYUL-X (ICLR, 2023).
Google DeepMind. OneTwo. GitHub https://github.com/google-deepmind/onetwo (2024).
Mallinar, N. et al. A scalable framework for evaluating health language models. npj Digit. Med. https://doi.org/10.1038/s41746-026-02492-x (2026).
Acknowledgements
We are grateful to the study participants who contributed their data to this research. We thank members of the consumer health research team at Google for feedback and technical support throughout this study (in particular S. Yuen, A. Pathak, J. Sunshine, F. Thng and J. Zhan); J. Shreibati and M. Thompson from the clinical team at Google for their feedback on the clinical utility and deployment of the proposed IR prediction model; the software development team who built the service that was used to recruit this large cohort and enabled remote collection of wearables and blood biomarker data (A. Dan, A. Badescu, D.-G. Stuparu, G.-I. Nitroi, S. Grigore, P. Navin and D. Trubnikov); our collaborators at Quest Diagnostics for their help in developing APIs to automate order placement and data retrieval; H. Maiwand for help creating the graphical abstract; the project management team (J. Galvan, L. Palao and S. Wemmer) for their efforts in coordinating the study and securing all necessary approvals; B. Winslow, N. Hammerquist, A. Mai, D. Peyton and E. Chung for help setting up the evaluation infrastructure for external endocrinologists to assess the IR agent; and T. Giest, H. Watkins, L. Cai, E. Blanchard, R. Luo, M. Liu, J. Gile and the entire human research laboratory at Google for their work in recruiting the validation cohort and collecting multimodal data.
Ethics declarations
Competing interests
This study was funded by Google. A.A.M., A.A.H., D.M., A.S., Z.E., A.Z.F., M.Z., X.L., Y.Y., M.M., C.H. and S.P. are employees of Alphabet and may own stock as part of the standard compensation package. G.N. and M.A.X. were interns at Google when the study was performed. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Christopher Hartshorn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Evaluation of IR prediction performance (classification).
a, Performance of our binary classification model for various input features for identifying insulin resistant individuals (using MAE + L1–L2 learners). Performance is evaluated using average area under the receiver operating characteristic curve (AUROC), sensitivity, specificity, precision, and area under the precision-recall curve (AUPRC). Error bars represent the standard deviation across the five folds. b, Visualization of the ROC curves for various feature sets across five cross-validation folds. Average values are colours, with the grey areas around each line indicating the standard deviation across the five folds. c, Visualization of the precision-recall curve for selected feature sets. Coloured lines represent the mean performance across fivefold cross-validation; shaded regions indicate standard deviation.
Extended Data Fig. 2 Interpretability and stratification.
a, Sankey diagram showing the relative feature importance (SHapley Additive exPlanations [SHAP] values) for each of the proposed nonlinear XGBoost models for direct regression. b,c, Qualitative evaluation of learned latent space’s interpretability of learning important features. The t-SNE reduced latent space shows that individuals with higher BMI (b) and RHR (c) are clustered closely together in space, following our quantitative results of classifying high BMI and high-RHR individuals using these learned representations. d,e, Distribution of individuals stratified by IR class and BMI classes (d) and IR versus physical activity classes as determined by number of daily steps (e). f, Results of classification performance for various lifestyle stratifications.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Metwally, A.A., Heydari, A.A., McDuff, D. et al. Insulin resistance prediction from wearables and routine blood biomarkers. Nature (2026). https://doi.org/10.1038/s41586-026-10179-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10179-2