Introduction
Asexual reproduction, including cloning, enables many species such as potatoes to thrive through efficient mass reproduction systems1,2. Even in animals, cloning from a body part, that is, producing offspring from somatic cells, is observed in a few lower species, such as pot worms and planaria3,4. However, higher animals, e.g., mammals, cannot reproduce clonally from somatic cells in nature, and the reason for this remains unknown5,6.
Until recently, mammalian cloning was not possible except in cases such as monozygotic twins7. However, following the development of nuclear transfer technology and the successful somatic cell cloning of frogs8, Dolly the cloned sheep9, and Cumulina the cloned mouse10, it has become possible to produce cloned animals even from terminally differentiated somatic cells11. This technology can be used to rescue endangered species12,13 or infertile individuals14 by increasing population numbers from somatic cells. Cloning is also thought to be useful for the large-scale production of superior-quality domesticated animals15,16, potentially making expensive meat, such as Wagyu beef, more affordable17,18.
However, mammalian cloning has not yet been put into practical use, not only due to its low cloning success rate, but also because of anomalies19 not observed in the cloning of plants or lower animals. These issues are believed to stem from failures in “reprogramming” the donor nucleus to reset the epigenetic state of differentiated somatic cells to that of a fertilized embryo; this occurs not directly due to DNA abnormalities, but the detailed mechanisms responsible remain unclear20.
If DNA abnormalities do not occur and epigenetic errors do not accumulate, mammals should be able to maintain their species through cloning. However, early studies showed that serial cloning in any mammalian species reached its limit after only a few generations21,22,23,24,25,26. This suggests that cloning or asexual reproduction may not be viable for maintaining mammalian species. Nevertheless, by improving cloning success using an epigenetic modification reagent that promotes nuclear reprogramming27, we succeeded in serially cloning up to 25 generations of mice28. Since the success rate improved slightly with each successive generation, we initially concluded that serial cloning could be continued indefinitely28. However, after continuing the experiment for 20 years, the success rate began to decline and eventually could not be sustained beyond 58 generations.
In this study, we aimed to investigate in detail what occurred in re-cloned mice after extended serial cloning, and to clarify why mammals maintain their species through sexual rather than asexual reproduction.
Results
Success rate of serial cloning up to 58 generations
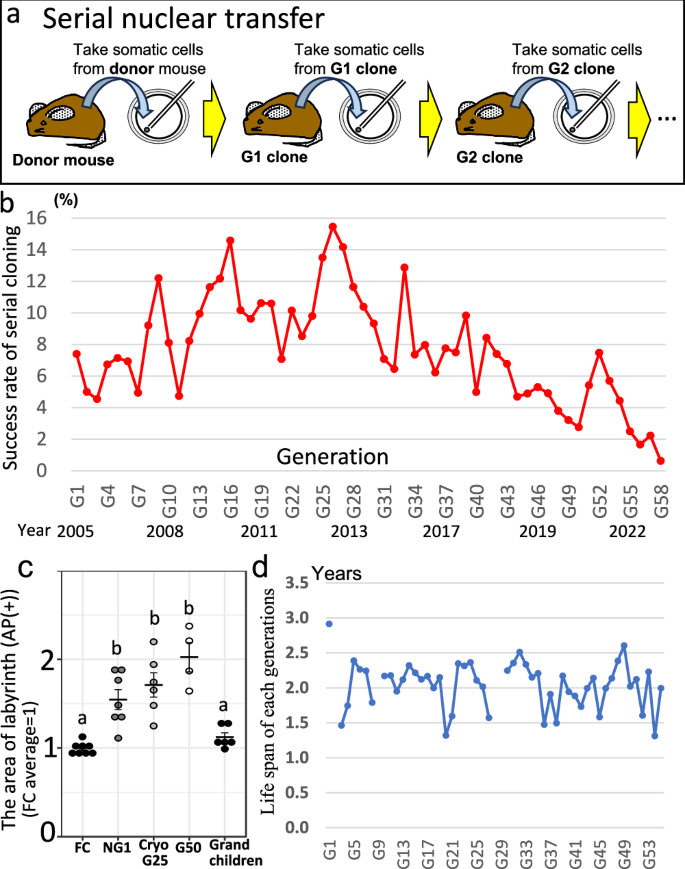
We began serial cloning experiments (Fig. 1a) in January 2005. The success rate of re-cloned mice gradually increased as generations progressed (up to 15.5% in the 26th-generation), and NGS analysis showed no difference between first-generation clones and 25th-generation re-clones. Therefore, we initially believed that serial cloning could probably continue indefinitely28, and thus continued this experiment. To maintain consistent experimental conditions as much as possible, TW and SW always participated in the production of next generation re-cloned mice. Approximately 3-4 generations of re-cloned mice were produced each year. However, the cloning success rate gradually decreased with each successive generation (Fig. 1b). Although success rates varied between generations, the average success rate declined to 0.6% after 57 rounds of serial cloning, and all 58th-generation re-cloned mice died the next day. In total, more than 1200 cloned mice were produced from a single original donor mouse (Supplementary Table 1). The original donor mouse was female with an agouti coat color; therefore, all cloned mice were female and exhibited the same coat color as the donor. Whole-genome sequencing (WGS) analysis also confirmed that these clones originated from the donor mouse (see later).
a Schematic showing the procedure for serial cloning multiple generations from the original donor mouse. A single donor was used to collect somatic cells (i.e., cumulus cells), which were then used for nuclear transfer to produce the first cloned mouse (“G1 clone”). When the G1 clone reached three months of age, cumulus cells were again collected to produce the next generation (G2 clone). This process was repeated over 20 years. b Success rate of mouse serial cloning across generations (n = 30,947 nuclear transfer). Data for G1–G25 were reported previously28. See also Supplementary Table 1. c Area comparison of placentas from re-cloned mice. The area of the labyrinth zone was measured at the center of the placenta. A value of 1 represents the mean area of the fertilized control. FC, fertilized control (n = 8, average 1 ± 0.06); NG1, newly generated clone (n = 7, average 1.55 ± 0.3); CryoG25, re-cloned mice derived from cryopreserved G24 cumulus cells (n = 6, average 1.71 ± 0.33); G50, re-cloned mice derived from G49 re-clones (n = 4, average 2.02 ± 0.33); Grandchildren, born from natural mating from offspring of G55 (n = 6, average 1.12 ± 0.12). Two-tailed unpaired Student’s t test. a vs. b: P < 0.002634–0.00871. See also Supplementary Fig. 1. d Lifespans of re-cloned mice from successive generations (n = 286 mice). For first-, second- and ninth-generation clones we could not measure mouse lifespans since these mice were used for other experiments. For clones from generations 28 and 29 we could not measure mouse lifespans due to reconstruction with our mouse facility. These mice were therefore sacrificed before they could die of old age. See also Supplementary Table 3. Source data are provided as a Source Data file.
Placenta and lifespan of re-cloned mice
All cloned mice have been found to possess abnormal placentas arising from epigenetic errors29,30. We therefore measured the body and placental weights of most cloned mice at the time of cesarean section. As controls, body and placental weights were also measured for mice derived from in vitro fertilized control embryos (FC), newly generated first-generation cloned mice (NG1), and 25th-generation re-cloned mice derived from the cryopreserved cumulus cells of G24 cloned mice (CryoG25). The mean body weight across re-cloned generations was ~1.88 g, slightly heavier than that of FC mice (1.3 g) but similar to other control clones (1.68–1.96 g) (Supplementary Table 2). The mean placental weight was about 0.30 g, which is two to three times heavier than that of FC mice (0.11 g), but comparable to other control clones (0.25–0.29 g). However, neither body nor placental weight increased over successive generations, suggesting that abnormalities did not accumulate. Histological examination and measurement of the area of labyrinth of these placentas showed slightly expanded the area of labyrinth zone when increased the number of serial cloning but no significant differences between NG1, CryoG25, and G50 cloned/re-cloned mice (Fig. 1c, Supplementary Fig. 1), despite expansion of the spongiotrophoblast layer, an abnormality specific to cloned mice31. However, these placentas were significantly different compared to those of FC mice. Finally, we also examined the average lifespan of randomly selected re-cloned mice across generations. The average was approximately two years (Fig. 1d, Supplementary Table 3), similar to that of the control cloned mice, suggesting no abnormality.
Effect of trichostatin A (TSA) on re-cloned mice
One possible explanation for the decreased success rate in later generations is that TSA, a known enhancer of nuclear reprogramming27, may lose its effectiveness in supporting the development of re-cloned embryos. When nuclear transfer was performed without TSA, the success rate for generating G51 clones was 1.6%, similar to that of NG1 (0.95%) and significantly lower than with TSA treatment (G51: 5.4%; NG1: 3.9%) (Fig. 2a, Supplementary Table 4). Thus, even after more than 50 nuclear transfer cycles, the somatic cell nuclei of G50 re-cloned mice retained sensitivity to TSA.
a Effect of TSA treatment on NG1 clones and G51 re-cloned mice. NG1 clones were derived from naturally conceived mice of the same genetic background, and G51 re-cloned mice were generated from G50 re-cloned mice. All experiments were performed with or without TSA. Statistical analysis was performed using a two-tailed, unpaired χ2-test; G51, n = 247 and 240, a vs. b: p = 0.0338; New G1, n = 330 and 333. a vs. b: p = 0.0026. See also Supplementary Table 4. b Immunostaining of one-cell stage zygotes of NG1 and G51using anti-H3K9me3 (top) or anti-acH3K14 (bottom) antibodies (red). Zygote nuclei were stained with DAPI (blue). Scale bar: 50 µm. c Relative fluorescence intensity of each pronucleus in FC zygotes and pseudo-pronucleus in cloned/re-cloned zygotes using anti-H3K4me3, H3K9me3, acH3K14, and H3K27me3 antibodies. The average signal in the female pronucleus of FC zygotes was standardized to 1. No significant differences were observed between NG1 and G51 for any epigenetic marks. Statistical analysis was performed using one-way ANOVA and Tukey-Kramer test (a vs b: p = 0.0350, n = 38 and 36, respectively. There was no difference between the other samples). See also Supplementary Fig. 2. d Embryo development rates to the blastocyst stage. 2C two-cell, 8C eight-cell, Mor morula, Bla blastocyst, Exp. B expanded blastocyst. Asterisks indicate significant differences between FC and NG1/G51 embryos. Statistical analysis was performed using a two-tailed, unpaired χ2-test; Morula, n = 126, 189 and 158, respectively. FC vs. NG1 or G51: P < 0.0001; Blastocyst, n = 115, 148 and 135, respectively. FC vs. NG1 or G51: P < 0.0001. Expanded Blastocyst, n = 91, 109 and 89, respectively. FC vs. NG1 or G51: P < 0.0001. See also Supplementary Table 5. e Immunostaining of blastocysts from FC (top) and G51 (bottom) embryos. Nuclei were stained with DAPI (top left, blue). Nanog-positive ICM cells appear red (top right), Cdx2-positive TE cells appear green (bottom left), and merged images are shown (bottom right). Scale bar: 50 µm. f Cell counts of TE (blue), ICM (orange), and double-negative cells (gray) in blastocysts from FC (n = 42), NG1 (n = 60), and G51 (n = 44) embryos. a vs. b: (p < 0.00001), c vs. d (p < 0.00001) and c vs. e (p = 0.00418) indicate significant differences. Statistical analysis was performed using a two-tailed, unpaired t test. See also Supplementary Table 6. Source data are provided as a Source Data file.
Epigenetic abnormalities in re-cloned zygotes produced by G50 cloned mice
We next examined whether the decreased cloning success was due to accumulated epigenetic abnormalities, as cloned embryos are known to differ epigenetically from fertilized embryos20,32,33. To investigate this, FC zygotes, NG1 clones, and G51 re-cloned zygotes were fixed and immunostained with antibodies targeting trimethylated histone H3 at lysines 4, 9, and 27 (H3K4me3, H3K9me3, H3K27me3), acetylated histone H3 at lysine 14 (acH3K14), and stained with DAPI. As shown in Fig. 2b, c, and Supplementary Fig. 2, the acetylation and methylation levels were similar between NG1 and G51 cloned/re-cloned zygotes, except for H3K9me3. However, both groups differed markedly from FC zygotes. These results suggest that epigenetic abnormalities did not accumulate in cloned/re-cloned embryos, at least at the one-cell stage.
In vitro development and quality of G51 cloned embryos
NG1 cloned embryos and G51 re-cloned embryos were cultured in vitro until they reached the blastocyst stage and were then compared with FC embryos. As shown in Fig. 2d and Supplementary Table 5, the developmental rate into expanded blastocysts was lower for NG1 and G51 embryos than for FC embryos. However, there were no differences between NG1 and G51 groups, suggesting that serial cloning up to 50 cycles did not impair in vitro embryo development.
To evaluate embryo quality, some blastocysts were analyzed for cell number and inner cell mass (ICM) allocation. ICM distribution did not differ clearly between groups (Fig. 2e, f, Supplementary Table 6). Although the average total cell number was significantly reduced in cloned/re-cloned embryos relative to FC embryos, no difference was observed between NG1 and G51 embryos. Interestingly, the average ICM cell number was slightly higher in cloned/re-cloned embryos compared to FC embryos.
RNA-seq analysis of G51 cloned blastocyst
To detect differences between FC embryos, NG1 clones, and G51 re-cloned embryos, we analyzed the whole-genome gene expression profiles of blastocysts using RNA-seq (Fig. 3a). Relatively well-developed blastocysts were selected, and RNA was extracted individually from each blastocyst. RNA-seq was performed, and the three groups were compared using principal component analysis (PCA) to identify gene expression differences. PCA showed that not only NG1 (green dots) and G51 (blue dots) clone/re-cloned embryos exhibited similar gene expression profiles, but these were also similar to those of FC (red dots) embryos.
a Gene expression profiles in blastocysts of FC, NG1, and G51 embryos. Principal component analysis (PCA), with PC1 and PC2 shown on the horizontal and vertical axes. Each dot represents an individual blastocyst. See also: accession number PRJDB37585. b Two pups derived from a G50 cloned mouse after natural mating with a male mouse. c Average litter size of FC mice (n = 6), G20 (n = 8), G50 (n = 16), and G55 re-cloned mice (n = 9) and their offspring after natural mating (n = 10). Statistical analysis was performed using a two-tailed, unpaired t test. a vs. b: p < 0.00001. See also Supplementary Table 7. d Histological examination of placentas derived from a G51 cloned mouse (top) and its grandchild (bottom). Scale bar, 1 mm. See also Supplementary Fig. 1. Source data are provided as a Source Data file.
Production of offspring from re-cloned mice by sexual reproduction
Although serial cloning could not continue beyond the 58th generation, the re-cloned mice remained healthy except G58, raising the possibility that subsequent generations could be produced via sexual reproduction. Therefore, FC mice, as well as G20, G50, and G55 re-cloned mice, were mated with normal male mice to evaluate pregnancy and offspring production. There was no difference in age at first delivery among the groups, and litter sizes were similar between G20 re-cloned mice (9.9) and FC mice (10.3). However, litter sizes declined sharply in G50 and G55 re-cloned mice (2.8 and 2.2, respectively) (Fig. 3b, c, Supplementary Table 7). The number of oocytes recovered from each generation was recorded when collecting cumulus cells as donor cells (Supplementary Table 8). Although superovulation was triggered by hormonal treatment, the number of oocytes did not decrease even with further generation of re-cloned mice. Therefore, the decrease in litter size was not due to a reduced number of oocytes ovulated from the re-cloned mice.
Interestingly, when the next generation (grandchildren) was produced from offspring of G55 re-cloned mice, the litter size increased to 7.0 (Fig. 3c). Moreover, histological analysis showed that placentas from the grandchildren were clearly smaller than the large placentas characteristic of cloned mice and were nearly the same size as those from fertilized embryos (Figs. 1c and 3d, Supplementary Fig. 1).
Normality of re-cloned mouse oocytes
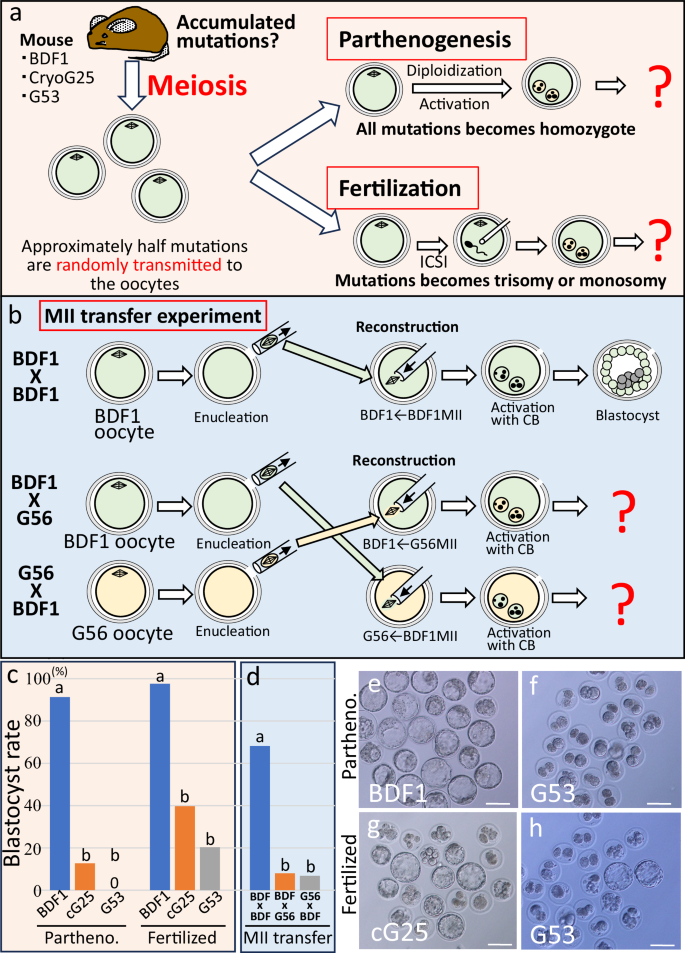
To examine whether oocytes derived from near-final-generation re-cloned mice were of reduced quality and/or prone to fertilization or developmental failure, we collected oocytes from CryoG25 and G53 re-cloned mice and subjected them to artificial activation and diploidization for parthenogenesis (Fig. 4a). If mutations accumulate through serial cloning, diploidization can reveal homozygous or defective mutations in oocytes, allowing assessment of their effects on early development. As a result, 91.3% of parthenogenetic embryos from BDF1 control mice developed into blastocysts, while only 12.7% of embryos from G25 re-cloned mice and none from G53 re-cloned mice developed into blastocysts (Fig. 4c, e, f, Supplementary Table 9).
a Schematic diagram of the oocyte quality test using BDF1, CryoG25 and G53 re-cloned mice. Approximately half of all mutations accumulated during serial cloning are randomly transmitted to oocytes. Some oocytes were used for diploid parthenogenesis (top), while others were fertilized via intracytoplasmic sperm injection (ICSI) (bottom), and blastocyst development rates were evaluated in vitro. b Schematic diagram of oocyte cytoplasm and MII quality test by replacement between G56 re-cloned mouse oocytes and fresh BDF1 oocytes. In the BDF1 x BDF1 control group (top), the metaphase II (MII) spindle was removed from a fresh BDF1 oocyte and immediately re-injected into the same oocyte, followed by parthenogenetic activation and diploidization. In the experimental group (bottom), the MII spindle was exchanged between a G56 re-cloned oocyte and a fresh BDF1 oocyte, followed by parthenogenetic activation and diploidization. c Developmental rate to the blastocyst stage of parthenogenetic embryos derived from BDF1 mice, CryoG25, and G53 re-cloned mice (n = 153, 57, 250, respectively) or fertilized embryos (n = 82, 77, 69, respectively). Statistical analysis was performed using a two-tailed, unpaired χ2-test. a vs. b: p < 0.0001. See also Supplementary Table 9. d Developmental rate to the blastocyst stage of parthenogenetic embryos derived from MII replacement oocytes between BDF1 and BDF1, BDF1 and G56, and G56 and BDF1. (n = 98, 79, 61, respectively). Statistical analysis was performed using a two-tailed, unpaired χ2-test. a vs. b: p < 0.0001. See also Supplementary Table 10. e Blastocysts derived from diploid parthenogenetic embryos of BDF1 mice (control). f Degenerated embryos derived from the diploid parthenogenetic embryos of G53 re-cloned mice. g Some blastocysts derived from fertilized embryos of CryoG25 re-cloned mice oocytes. h A few blastocysts derived from fertilized embryos of G53 re-cloned mice oocytes. e–h Scale bar: 100 µm. Source data are provided as a Source Data file.
Next, oocytes from re-cloned mice were fertilized in vitro with sperm from normal males (Fig. 4a). Even if oocytes carried many lethal mutations, normal sperm genes could improve blastocyst development. As a result, 97.6% of BDF1 control embryos reached the blastocyst stage, compared with 39.7% from G25 re-cloned mice and 20.3% from G53 re-cloned mice (Fig. 4c, g, h, Supplementary Table 9). These results clearly indicate that the number of lethal mutations in oocytes increases with each generation of serial cloning.
Finally, to determine whether the abnormalities originated from the oocyte nucleus (Metaphase II; MII) or cytoplasm, we performed MII replacement between oocytes from G56 re-cloned mice and fresh BDF1 mice (Fig. 4b). When the MII from one BDF1 oocyte was removed and immediately transferred into same oocyte, 68.2% of parthenogenetic embryos developed into blastocysts, confirming normality of both nucleus and cytoplasm (Supplementary Table 10). When MII from G56 re-cloned oocytes was transferred into enucleated BDF1 oocyte cytoplasm, only 8.0% of embryos developed to blastocysts. Similarly, when BDF1 MII was transferred into enucleated G56 oocyte cytoplasm, blastocyst formation was only 6.8%. These results suggest that G56 oocytes are damaged in both nucleus and cytoplasm.
Whole-genome sequencing (WGS) analysis of re-cloned mice
To investigate genomic mutations accumulated across generations, we conducted WGS on re-cloned mice from various generations. Short-read NGS-based WGS (read coverage: >30×) was performed on ten individuals from generations G6 to G57 (Fig. 5a, Supplementary Table 11). In addition, three of these individuals (G6-2, G50-1, and G57-2) underwent long-read WGS (read coverage: >20×) using Oxford Nanopore Technology long-read data.
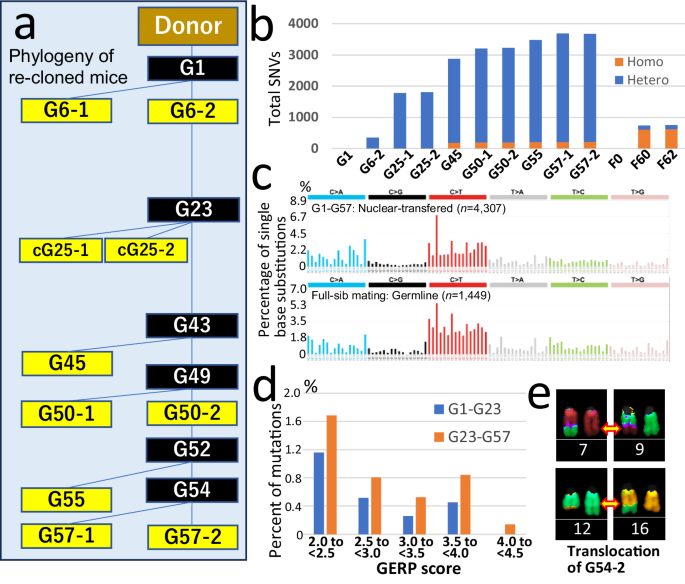
a Phylogenetic tree of re-cloned mice. Yellow boxes indicate individuals analyzed by WGS. Black boxes indicate generations of common ancestors estimated from WGS data. b Total number of de novo SNVs detected in each re-cloned mouse (G1 to G57-2) and in each mouse of sib-mating mouse lines (F0–F62) as per WGS analysis. Blue bars represent heterozygous variants; orange bars represent homozygous variants. See also Table 1. c Top: mutation signatures accumulated from G1 to G57. Bottom: mutation signatures observed in sibling breeding lines of wild-type mice from F0 to F62. See also Supplementary Fig. 3. d Proportion of mutations with a GERP score ≥2, indicating evolutionarily conserved regions, divided between generations G1–G23 (blue) and G23–G57 (orange). See also Supplementary Table 14. e G54-2 re-cloned mouse exhibited two large-scale chromosomal translocations: between Chr 7 and Chr 9 (top), and between Chr 12 and Chr 16 (bottom), as demonstrated by SKY–FISH chromosome painting. See also Supplementary Fig. 4. Source data are provided as a Supplementary Data 1.
Based on single nucleotide variant (SNV) profiles, we reconstructed the genetic lineage of each individual. As shown in Fig. 5a, yellow boxes indicate individuals analyzed by WGS, while black boxes denote estimated common ancestors. This phylogeny matched the experimental lineage, confirming that no genetic contamination occurred during the 20-year serial cloning process. SNVs accumulated over generations (Fig. 5b), with ~3700 SNVs and 80 insertion/deletions (indels; excluding repeat elements) identified from G1 to G57 (Table 1). On average, each generation acquired 69.4 SNVs and 1.4 indels. The SNV mutation signatures from G1 to G57 resembled those of naturally occurring germline mutations34 (Fig. 5c, Supplementary Fig. 3).
From G1 to G57, we identified 80–84 de novo structural variants (SVs) (Supplementary Data 1). Table 2 lists 16 large SVs (LSVs) of ≥40 kb, which are more likely to affect phenotypes. Many SVs appeared between G25 and G45. Among them, loss of one X chromosome, loss of heterozygosity (LOH) on chromosome 4 (147.5 Mb), and a translocation between chromosomes 7 and 9 are likely highly deleterious for fitness. LSVs also occurred after G45. Chromosomal abnormalities, including the X chromosome loss and two translocations, were likewise confirmed in the karyotype analysis of G54 re-cloned mice (Fig. 5e, Supplementary Fig. 4). We measured telomere length using short-read WGS and Telomerecat software 4.0.0. Although telomere length estimates varied widely and were method-dependent across samples, significant telomere shortening between G6 and G55 is unlikely (Supplementary Table 12). Nanopore long-read WGS data also revealed no evidence of extreme telomere shortening at individual chromosome ends (≥10 kb in G57-2).
To evaluate the effects of accumulated mutations, we analyzed their effects on genes. Of the 4404 SNVs and small indels identified, 0.95% were classified by SnpEff as “High” (loss-of-function) or “Moderate” (mainly missense). Dividing mutations into two groups—those from G1 to G23 during increased cloning success and those from G23 to G57 during decreased success (Fig. 1b)—revealed a significantly higher proportion in the latter group (G1–G23: 0.52%; G23–G57: 1.19%; P = 0.034) (Supplementary Table 13). Consistent with this, we assessed potentially deleterious variants using Genome Evolutionary Rate Profiling (GERP) scores, which evaluate functional constraint across coding and noncoding regions. The proportion with GERP scores ≥2 was significantly higher in the latter group (G1–G23: 2.4%; G23–G57: 4.0%; P = 0.005, Fisher’s exact test) (Fig. 5d, Supplementary Data 1, Supplementary Table 14). Mutation spectra also shifted, with transversions rising from 44.0% in G1–G23 to 47.9% in G23–G57 (P = 0.015, Fisher’s exact test) (Supplementary Fig. 3). In terms of SVs, G57 re-cloned mice showed at least 16 variants affecting exons (Supplementary Data 1). Across generations, two large deletions were identified in an adjacent region, resulting in homozygous loss of Csmd1 gene function (Supplementary Fig. 5). Considering the complete mutation spectrum, ~30 loss-of-function mutations and ~50 missense mutations had accumulated in G57 mice, even when excluding X chromosome loss (Table 1). In this context of an extensive background mutation load, the emergence of additional deleterious mutations near the final generation, including Csmd1 disruption, may have exceeded a tolerable threshold, leading to lethality or severe developmental impairment, ultimately contributing to the collapse of cloning success (Supplementary Fig. 6). Considered together, these findings indicate that the progressive decline of serial cloning success is strongly associated with the accumulation of de novo mutations—including large SVs and harmful SNVs/indels after G23–G25—rather than the effect of any single mutation alone.
To compare mutation patterns in asexual versus sexual reproduction, we analyzed WGS data from two mouse lines subjected to 60 generations of sibling mating over ~20 years. These lines accumulated 739 and 752 SNVs, 22 and 23 INDELs and 29 and 29 SVs, respectively, with ~80% of autosomal variants being homozygous (Fig. 5b, Table 1). This corresponds to 22.3 SNVs, 0.68 INDELs and 0.91 SVs per generation, representing one-third of the SNV mutation rate in serial cloning and a slightly lower INDELs and SV occurrence frequency.
Finally, we performed WGS again on offspring produced via natural mating between G55 re-cloned mice and wild-type males (C57BL/6), as well as on their grandchildren. As expected from Mendelian inheritance, the mutations present in G55 were passed to both generations. LSVs transmitted across generations are listed in Supplementary Table 15. Notably, the translocation between chromosomes 7 and 9 in G55 was also found in G55 descendants, likely because both affected chromosomes were inherited together, preserving balance.
Discussion
It has long been unclear whether mammals, unlike plants and some lower animals, could sustain their species through clonal reproduction alone. In this study, we performed continuous somatic cell nuclear transfer cloning in mice for nearly two decades, covering 57 generations. Our findings show that clone-specific epigenetic abnormalities did not accumulate, but genomic mutations did, increasing the burden of deleterious variants. Ultimately, this accumulation drove a progressive decline in birth rates, making continued clonal reproduction impossible.
On average, each generation acquired 70 single-nucleotide variants (SNVs) and 1.5 structural variants (SVs), including large structural variants (LSVs). These mutation rates are comparable to those in certain mouse somatic cell lineages35,36 and in the human germline37, suggesting that the rate itself is not unusually high. Nonetheless, two factors appear central to the collapse of long-term clonal propagation: (1) the emergence of LSVs, which are rarely seen in the germline37,38, and (2) the absence of chromosomal recombination in clonal reproduction, which prevents effective purifying selection and allows all classes of mutations to accumulate progressively.
A critical turning point occurred around generation G25. Before this stage, chromosomal abnormalities and coding mutations were efficiently eliminated, and birth rates even improved. During this period, although we were not able to detect adaptive mutations by WGS analysis, some mutations may have transiently enhanced cloning performance, or somatic cells that were more easily reprogrammable and capable of generating viable re-clones may have been preferentially selected, resulting in a positive selection effect. However, these advantages were not sustained. After G25, large and deleterious SVs, including loss of heterozygosity (LOH) and X chromosome loss, became prominent. The build-up of harmful variants appears to have outweighed adaptive effects, such as the positive selection effect around G25, resulting an irreversible decline in birth rates. The frequency of deleterious mutations nearly doubled, from 0.86 to 1.78 per generation (Table 1), in parallel with the onset of this irreversible decline. This transition suggests that the burden of accumulating mutations began to affect not only nuclear genomic integrity but also oocyte-level functions essential for development. Consistent with this interpretation, our MII replacement experiments indicate that late-generation oocytes exhibited compromised cytoplasmic competence in addition to nuclear defects.
This study was conducted over a period of ~20 years. Therefore, considering variations to the quality of culture medium or equipment, it was impossible to maintain completely identical experimental conditions throughout the entire period. However, two researchers (TW and SW) consistently participated in the nuclear transfer experiments across all generations. Furthermore, nuclear transfer experiments conducted during this period using BDF1 cumulus cells demonstrated no change to the birth rate of cloned mice, including the duration of the COVID-19 pandemic (Supplementary Table 16). Therefore, it is unlikely that these factors contributed to the decreased cloning success rate. Conversely, in this study, TSA treatment was conducted during nuclear transfer for production of all generations of re-cloned mice. TSA is a standard class I/II histone deacetylase (HDAC) inhibitor known to not only activate genes involved in transcription and promote nuclear reprogramming27,39, but also reactivate endogenous retroviruses40,41. However, since a very low concentration (50 nM) was used during nuclear transfer and lasted only about 8 h, it is unlikely that TSA treatment caused the high mutation rate of the re-cloned mice.
Late-generation clones except last generation that survived to birth were notably healthy, with normal lifespans, despite carrying numerous deleterious mutations. Although placental size tended to increase with advancing generations, embryonic development itself was similar between early and late groups. Likely, re-cloned embryos that acquired lethal mutations were lost after implantation, and only those without such mutations developed to term. This stringent selection probably ensured that surviving re-cloned mice remained healthy. WGS analysis was performed using samples collected from live-born re-cloned mice, so it may not have identify critical mutations that cause lethality in embryos or fetuses.
Even more striking was the preservation of reproductive capacity. Despite over 20 years of repeated clonal propagation without engagement of the germline or reproductive organs, all examined re-cloned mice retained fertility. Since sexual system function is not required for the survival of individual clones, it is improbable that only cloned embryos with normal reproductive system were selected and born. Instead, these results imply that the sexual reproductive system is unusually robust against mutational damage, even under conditions that foster accumulation of deleterious variants. Throughout this study, we consistently utilized oocytes collected from young individuals, and it is known that the oocytes of young individuals can effectively suppress age-related DNA damage42, and this ability may have been particularly effective against mutations in the sexual reproductive system.
The necessity of sexual reproduction has traditionally been explained by the Red Queen hypothesis, which posits that sexual reproduction is required to maintain genetic diversity and thereby enable species to persist under changing environmental conditions6,43. Our study, however, reveals limitations of this framework. Re-cloned mice displayed nearly threefold higher mutation rates than their sexually reproducing counterparts, thus generating diversity even without recombination. Moreover, because our experiments were performed under specific-pathogen-free conditions with minimal environmental pressure, the failure of clonal reproduction in this setting cannot be attributed to the Red Queen hypothesis.
In contrast, our results align closely with Muller’s ratchet theory44. This model predicts that in asexual lineages, deleterious mutations inevitably accumulate, ultimately producing mutational meltdown and extinction45. Across more than 30,000 nuclear transfer attempts spanning 58 generations, our study provides the first empirical demonstration that mutational meltdown, as predicted by Muller’s ratchet, can indeed occur in mammals. In comparison, wild-type mice maintained through natural breeding showed no fertility decline across more than 60 generations, emphasizing that the mutational load of clonal reproduction exceeds the threshold compatible with long-term mammalian survival. At the same time, the observation that most re-cloned mice could remain healthy despite harboring numerous genetic alterations offers insight into the extent of mutation tolerance at the level of individual organisms.
Since the birth of Dolly the sheep9, cloning technology has been envisioned for diverse applications, including agricultural improvement, regenerative medicine, and the conservation of Earth’s rare genetic resources46,47,48. Yet, these findings demonstrate that practical application of cloning of mammals requires a deeper appreciation of these biological constraints, warranting further research, and reaffirm the evolutionary inevitability that sexual reproduction is indispensable for the long-term survival of mammalian species.
Methods
Animals
A three-month old BD129F1 (BDF1 × 129/Sv) mouse was used as the original donor animal, and cumulus cells were used as nuclear donors. This mouse was produced in our laboratory using three-way crosses between the C57BL/6, DBA/2, and 129/Sv strains. During experimentation, C57BL/6J mice were maintained at a constant temperature (25 °C ± 1 °C) under a 12-h light/dark cycle, with ad libitum access to food and water. The donor mouse that showed the highest success rate in producing first-generation cloned mice (hereafter, “G1”) was selected as the original donor and was used to initiate the serial mouse cloning experiment.
To generate subsequent generations, recipient oocytes were collected from adult BDF1 females. The surrogate pseudopregnant females used as embryo transfer recipients were ICR strain mice mated with vasectomized males of the same strain. BDF1, C57BL/6J, and ICR mice were purchased from Shizuoka Laboratory Animal Center (Hamamatsu, Japan).
For all sexual reproduction experiments, animal protocols were approved by the Animal Committee of Osaka University (Osaka, Japan; Approval No. FBS-17-006) and the Experimental Animal Care Committee of the Radiation Effects Research Foundation (RERF; Hiroshima, Japan; Approval No. 2018-02). All experiments were conducted in accordance with the 1996 National Institutes of Health guidelines and the Guidelines for Animal Experiments of the RERF. During experimentation, C57BL/6J mice were maintained at a constant temperature (22 °C ± 1 °C) under a 12-h light/dark cycle, with ad libitum access to food and water.
Housing conditions and diet were kept constant throughout the experiment. On the day of experimentation or after completion of all procedures, mice were euthanized by CO₂ inhalation or cervical dislocation. All animal experiments followed the ARRIVE guidelines and were performed in accordance with the Guidelines for the Care and Use of Laboratory Animals, with approval from the Institutional Committee of Laboratory Animal Experimentation of the University of Yamanashi (Reference No. A29-24).
Oocyte collection
Female mice were superovulated by injection of 5 IU of equine chorionic gonadotropin, followed 48 h later by 5 IU of human chorionic gonadotropin (hCG). Cumulus–oocyte complexes (COCs) were collected from the oviducts 14–16 h later and transferred to a Falcon dish containing HEPES–CZB medium49. To disperse the cumulus cells, COCs were placed in a 50 μL droplet of HEPES–CZB medium containing 0.1% bovine testicular hyaluronidase for 3 min. Cumulus-free oocytes were washed twice and transferred to 20 μL droplets of CZB medium50 for culturing.
Collection of cumulus cells and tail tip fibroblasts
Cumulus cells were collected from donor or re-cloned mice using the same method as for oocyte collection. Fibroblasts were collected from the tail tips of G54 re-cloned mice for karyotype analysis as described previously51. Briefly, tail tips were skinned, cut into small pieces, then incubated in 5 mL DMEM (Sigma-Aldrich, St. Louis, MO, USA) supplemented with 10% fetal calf serum (Sigma-Aldrich). After 10–14 days at 37.5 °C in 5% CO₂ in air, proliferating fibroblasts were dissociated using trypsin and replated into larger dishes to expand the cell population.
Nuclear transfer
For nuclear transfer, oocytes were first transferred to a droplet (~10 μL) of HEPES–CZB containing 5 μg/mL cytochalasin B, placed under oil on the microscope stage. Each oocyte was held with an oocyte-holding pipette, and its zona pellucida was perforated by several piezoelectric pulses applied to the tip of an enucleation pipette10. The metaphase II chromosome spindle complex, visible as a translucent spot in the ooplasm, was drawn into the pipette with a small amount of ooplasm and gently pulled away until the cytoplasmic bridge was pinched off. Enucleated oocytes were then injected with a cumulus cell nucleus recovered from a 3-month-old re-cloned mouse, as described previously10. Briefly, the zona pellucida and the oocyte membrane were perforated by piezoelectric pulses, and the donor nucleus was inserted into the ooplasm. Reconstructed oocytes were transferred into CZB medium and incubated at 37 °C in 5% CO₂.
Effects of nuclear transfer techniques on birth rate
During this long-term study, we improved the birth rate of cloned mice by optimizing nuclear transfer methods, such as using different histone deacetylase inhibitor (HDACi) treatments instead of TSA52,53. However, to isolate the effect of re-cloning, the standard method described above was used whenever possible. Some variations, such as in media quality or experimenter skill, were unavoidable over the study period and may have affected success rates or phenotypes. To control for these factors, we generated cloned mice from other BD129F1 donors as technical or time-matched comparisons (designated NG1). To assess the pure effect of serial nuclear transfer, cumulus cells from G24 re-cloned mice were frozen in June 2013 and thawed 8 years later to generate additional G25 mice (designated CryoG25), concurrently with G50 generation. Fertilized control (FC) mice with the same BD129F1 genetic background were also produced via intracytoplasmic sperm injection (ICSI), simulating equivalent in vitro manipulation and culture stress.
Activation of cloned embryos
Reconstructed oocytes were activated using 5 mM SrCl₂ in Ca²⁺-free CZB medium containing 50 nM trichostatin A (TSA) and 5 μM latrunculin A for 9 h27,33,54. Pseudo-pronuclear formation was verified, and embryos were cultured for either 1 day before transfer or 96 h to evaluate blastocyst development.
Embryo transfer
On the day following mating with vasectomized males, 2-cell stage cloned embryos were transferred into the oviducts of pseudo-pregnant ICR females at 0.5 days post coitum (dpc)55. On the transfer day, recipients were anesthetized by intraperitoneal injection of medetomidine, midazolam, and butorphanol. Five to eight embryos were transferred per uterine horn, and atipamezole was administered. At 19.5dpc, offspring were delivered by cesarean section and transferred to foster mothers that had delivered naturally. Fertility was assessed by mating offspring at three weeks of age.
Intracytoplasmic sperm injection (ICSI)
To generate genetically identical but fertilized control (FC) mice, oocytes were collected from BDF1 females and sperm from 129/Sv males, producing BD129F1 offspring as with the original donor. To match cloned embryo conditions, FC mice were generated via ICSI, with equivalent in vitro manipulation and 1 day of CO₂ incubation49.
Histological analysis of placentae
Placentae from FC, NG1 (newly generated clone), G25, G50 re-cloned mice, and grandchild mice were embedded in OCT compound (Sakura Finetek, Japan). Blocks were cryosectioned at 7 μm (Thermo Fisher Scientific, USA) and mounted on Superfrost Micro Slides (Matsunami Glass, Japan). Sections were fixed in 4% paraformaldehyde for 10 min at room temperature, washed three times in PBS, incubated in 100 mM Tris-HCl (pH 9.5) for 10 min, and stained using NBT-BCIP (Vector Laboratories) for endogenous alkaline phosphatase (AP) activity. Sections were rinsed in Tris-HCl and counterstained with Nuclear Fast Red (Vector Laboratories). Trophoblasts in the labyrinth layers were detected by AP staining, and the area of AP staining-positive labyrinth layers were compared between natural, re-cloned mice and grandchild mice.
Immunostaining of re-cloned embryos at the one-cell stage
Ten hours post-activation, oocytes were fixed for 20 min at 25 °C in 4% (w/v) paraformaldehyde with 0.2% (v/v) Triton X-100 (Nacalai Tesque, Kyoto, Japan). Fixed oocytes were washed thrice in PBS with 0.25% BSA (Millipore, Darmstadt, Germany) and stored at 4 °C. Embryos were incubated with primary antibodies: rabbit anti-H3K4me3 (1:1000, Millipore, Darmstadt, Germany), rabbit anti-H3K9me3 (1:1000,Millipore, Darmstadt, Germany), or rabbit anti-H3K27me3 (1:1000, Millipore, Darmstadt, Germany) in PBS-BSA overnight at 4 °C. Antibodies were supplemented with 0.2% (v/v) Tween (Millipore, Darmstadt, Germany). After two PBS-BSA washes (15 min each), embryos were incubated for 45 min with Alexa Fluor Cy5-labeled goat anti-rabbit IgG (1:500, Abcam, Cambridge, UK). DNA was stained with 2 μg/mL DAPI (Molecular Probes Inc., OR, USA). The largest pseudo-pronucleus per embryo was imaged using confocal microscopy (FV-1000, Olympus Corp., Tokyo, Japan). Fluorescence intensity was quantified using Olympus Fluor View and NIH ImageJ.
Blastocyst immunostaining
To assess cloned blastocyst quality, cell counts were performed via immunofluorescence staining as described previously56. Briefly, blastocysts were fixed for 20 min at 25 °C in 4% paraformaldehyde with 0.2% Triton X-100, washed thrice in PBS with 0.25% BSA, and stored at 4 °C. Primary antibodies: anti-CDX2 mouse monoclonal (1:500, BioGenex, CA, USA) for TE cells and anti-Nanog rabbit polyclonal (1:500, Abcam, Cambridge, UK) for ICM cells. Secondary antibodies: Alexa Fluor 488-labeled goat anti-mouse IgG and Alexa Fluor 568-labeled goat anti-rabbit IgG (1:500, Abcam, Cambridge, UK). DNA was stained with 2 μg/mL DAPI (Molecular Probes, OR, USA). Blastocysts were mounted on slides, and serial z-stacks were captured by confocal microscopy (FV-1200). Cell numbers were quantified using Olympus Fluor View.
Re-cloned mice oocyte quality examination
Oocytes were collected from BDF1, G25, and G53 re-cloned mice, and diploid parthenogenetic or fertilized embryos were generated to assess blastocyst development. Diploid parthenogenetic embryos were produced using the same method as that used to activate reconstructed oocytes after nuclear transfer. ICSI was performed using the same procedure as that for generating FC mice. The resulting parthenogenetic and fertilized embryos were cultured for five days, and their development to expanded blastocysts was evaluated. Next, to investigate the nucleus (metaphase II: MII) and cytoplasm of cloned mice oocytes separately, we performed MII replacement between oocytes from BDF1 and G56 re-cloned mice. This was followed by diploid parthenogenesis treatment to determine the rate of development to the blastocyst stage. MII extraction was performed using the same method as described for oocyte enucleation. However, when transferring MII to enucleated oocytes, the enucleation pipette was used to inject the MII into enucleated oocytes in cytochalasin B-containing medium, after which the hole was immediately sucked out to seal the cell membrane57. After replacing the MII, all oocytes were allowed to rest in an incubator for at least two hours before activation.
RNA-seq of re-cloned blastocysts
Five blastocysts were sampled from each of the following groups: FC mice, NG1 cloned mice, and G50 re-cloned mice. Samples were frozen immediately in liquid nitrogen and stored at –80 °C until further analysis. RNA-seq libraries were prepared using the SMART-seq Stranded kit (Takara Bio, Shiga, Japan) and sequenced as 150-base single-end reads on the DNBSEQ-G50RS system (MGI Tech., Shenzhen, China). Sequence data were then mapped against the mouse reference genome sequence (GRCm39/mm39) using STAR58. We then calculated the “transcripts per million” value for each gene using RSEM59. To extract transcripts that often exhibited upregulation in blastocysts greater than twofold, we then calculated the fold change of each signal in all individuals within each group (i.e., NG1 and G50). These were then compared with the mean signal in the FC; the mean difference was then assessed using the DESeq2 R package60.
WGS and mapping
Genomic DNA was extracted from spleen samples using smart DNA prep (m) (Analytik Jena) and Qiagen Genomic tip DNA extraction kits (QIAGEN). For short-read WGS, paired-end libraries were prepared with DNA samples without PCR amplification using a TruSeq DNA PCR-Free Library Prep Kit (Illumina). All libraries were then sequenced on NovaSeq 6000 or X plus platforms (Illumina) to obtain paired-end 151-bp reads. Short-read sequence data were then mapped to the mouse reference genome (UCSC mm10) using BWA-MEM version 0.7.1761 with the “-M” option. This mapping was also processed with Picard version 2.18.26 (broadinstitute.github.io/picard) to remove PCR duplicates. In the analysis of sib-mating mouse lines, we utilized the ancestral genome data (F0 mice) from our previous paper (Ancestor ConA2 male-female mice). For long-read sequencing, libraries were prepared from G6-2, G50-1, and G57-2 using the SQK-LSK114 kit (Oxford Nanopore) and sequenced on a PromethION with FLO-PRO114M flow cells in high-accuracy mode. Base calling was performed with Guppy version 6.4.2 and Dorado version 7.2 (https://community.nanoporetech.com/). Base-called reads were mapped to the mouse reference genome (UCSC mm10, excluding chrY) using minimap2 version 2.21 and version 2.2862 with the parameters “-ax map-ont --MD.”
Identification of de novo mutations
Genomic variants were called from short-read bam files using GATK HaplotypeCaller version 4.1.0.063 with a minimum base quality of 20. For accurate detection of somatic mutations, effective whole-genome coverage (EWC) regions were defined per sample based on the following criteria: (i) MQ60 read depths were between 50% and 300% of the chromosome-specific peak coverage; (ii) MQ60 reads comprised at least 80% of mapped reads at each site; and (iii) a minimum base quality of 20 was confirmed using SAMtools version 1.964. Only shared EWC regions were used for mutation analyses (excluding SVs), covering 76.4% of the diploid genome for serial nuclear transfer lines, and 78.6% for sib-mating mouse lines (Supplementary Table 11). All de novo mutations were identified from filtered variants using the following criteria: (1) located within EWC regions; (2) multiallelic variants excluded; (3) variants with 0.35 > AF ≥ 0.03 in G45, G57-1, and G57-2 but <0.03 in others were excluded to remove NovaSeq X Plus–specific noise; (4) minimum VAF = 0 in any clone; (5) Fisher strand bias (FS) < 23; (6) variants with AF ≥ 0.06 and <0.2 detected in ≥3 samples excluded; (7) max VAF ≥ 0.3 for SNVs and ≥ 0.25 for INDELs in G6, G25, G45, G50, G55 and G57. We also used the following INDEL-specific filters: (8) exclude variants in homopolymers (>3 bp) or short repeats (e.g., di-, tri-, tetra-, penta-, or hexa-). (9) Only manually confirmed variants in IGV were retained. (10) Finally, we retained variants that met either of the following: (i) VAF < 0.10 in G6-1; or (ii) VAF < 0.10 in G6-2 and, in G6-1, VAF ≥ 0.30 for SNVs or VAF ≥ 0.25 for INDELs. In the analysis of sib-mating mouse lines, we retained variants that satisfied the following: VAF < 0.06 in both the F0 male and F0 female, and—on autosomes—VAF ≥ 0.30 for SNVs or ≥0.25 for INDELs in F60 or F62; on sex chromosomes, VAF ≥ 0.80 in F60 or F62.
SV calling using short- and long-read data
SV calling of short-read data was conducted using Manta version 1.6.065, RUFUS version 1.066, and Delly version 0.8.667. SV calling by Manta was conducted using the following criteria: 1) The number of split reads for the “ALT” allele (SRA) was >3 in a mouse. 2) The minimum SRA number of other mice was zero. SV calling by RUFUS was conducted using the following criteria: 1) “SVTYPE” was described in the INFO column of the VCF file, or 2) the length of the “REF” allele or the “ALT” allele was >40 bp. SV calling by DELLY was conducted using the following criteria: 1) In all cases the number of variant reads (i.e., DV or RV) was >3 for a mouse. 2) The minimum number of variant reads in other mice was 0.
Next, we performed SV detection using long-read data to recover mutations missed in short read analyses. Mutation candidates were then extracted from G6-2 and G50-1 using sniffles version 1.0.1268 with the parameter: “-s 3.” VCF files of SV calls from each sample were merged using SURVIVOR version 1.0.769 with the following parameters: max distance between breakpoints was 1000 bp and “types must match.” The merged VCF file was then used to perform a “force call” of SVs using sniffles. Force-called VCF files of each sample were again merged using SURVIVOR. Structural variants were called using the following inclusion criteria: (1) SVTYPE is TRA or SVLEN is ≤−40 or ≥40, (2) The number of reads supporting alt is 0 for G6-2, >5 for G50-1, and/or G57-2, and (3) depth is >10 and <100 for both G6-2 and G50-1. All candidates were then manually inspected by IGV, including short read results. Using this procedure five new de novo genes were detected. Three of these could be linked to the generation in which they arose using short read sample information. Finally, we note that this analysis was not limited to EWC regions.
Mutation analysis
The genetic lineage of each re-cloned mouse was inferred from its SNV profile. Assuming a constant number of SNVs per generation, lineage divergence points were identified. By excluding regions affected by LOH, large deletions, or X chromosome loss, we constructed a consistent phylogenetic relationship.
Mutational signature analysis and visualization of single-base substitutions were carried out using SigProfilerAssignment70 (https://cancer.sanger.ac.uk/signatures/assignment/app/), with the parameters set as follows: Reference genome = “mm10/GRCm38,” Variant class = “SBS.” Reference mutational signatures = “COSMIC v3.4.” Type of study = “Whole-Genome Sequencing”70. Functional annotations were performed with SnpEff version 4.3t71. De novo mutations were further annotated using GERP scores72. GERP score data derived from the mouse mm9 reference was obtained from the UCSC website (http://hgdownload.soe.ucsc.edu/gbdb/mm9/bbi/All_mm9_RS.bw).
Telomere length evaluation
Telomere length was assessed using WGS data from two G6 individuals, two CryoG25 individuals, and two G50 individuals by Telomerecat (version 4.0.0)73 using the default parameters. Although we performed whole-genome sequencing on either the Illumina NovaSeq 6000 or NovaSeq X Plus platforms, only datasets produced on the NovaSeq 6000 were used for telomere length analysis in order to ensure methodological consistency and avoid potential platform-specific biases.
Karyotyping of G54 re-cloned mice
Chromosomes from tail-tip cells of two G54 re-cloned mice (G54-1 and G54-2) were analyzed by spectral karyotyping using fluorescence in situ hybridization (SKY–FISH) chromosome painting techniques (SCAS Ltd., Tokyo, Japan).
Statistical analyses
Most serial cloning experiments for each generation were repeated at least twice. Other experiments were independently performed at least three times by two to three experimentalists, and similar results were obtained regardless of the experimenter. Rates of embryo development, offspring birth, and body and placental weights were evaluated using chi-squared tests. Fluorescence intensity was analyzed by one-way ANOVA followed by Tukey’s multiple comparison tests as implemented in GraphPad Prism (GraphPad Software, USA). Principal component analysis (PCA) was performed using the DESeq2 R package. For all statistical tests, a value of P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available in the main text, Supplementary Figs./tables, or source data file. NGS and WGS data have been deposited in DNA Data Bank of Japan Sequence Read Archive under accession numbers PRJDB37585 and PRJDB35828. Additional data related to this paper may be requested from the authors. Source data are provided with this paper.
References
Gerritsen, J. Sex and parthenogenesis in sparse populations. Am. Naturalist 114, 718–742 (1980).
Lively, C. M. & Morran, L. T. The ecology of sexual reproduction. J. Evol. Biol. 27, 1292–1303 (2014).
Yoshida-Noro, C. & Tochinai, S. Stem cell system in asexual and sexual reproduction of Enchytraeus japonensis (Oligochaeta, Annelida). Dev. Growth Differ. 52, 43–55 (2010).
Nishimura, O. et al. Unusually large number of mutations in asexually reproducing clonal planarian Dugesia japonica. PLoS One 10, e0143525 (2015).
Avise, J. C. Evolutionary perspectives on clonal reproduction in vertebrate animals. Proc. Natl. Acad. Sci. USA 112, 8867–8873 (2015).
West, L. R. A pluralist approach to sex and recombination. J. Evolut. Biol. 12, 1003–1012 (1999).
Blickstein, I. & Keith, L. G. On the possible cause of monozygotic twinning: lessons from the 9-banded armadillo and from assisted reproduction. Twin Res. Hum. Genet. 10, 394–399 (2007).
Gurdon, J. B. Adult frogs derived from the nuclei of single somatic cells. Dev. Biol. 4, 256–273 (1962).
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J. & Campbell, K. H. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Wakayama, T., Perry, A. C., Zuccotti, M., Johnson, K. R. & Yanagimachi, R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394, 369–374 (1998).
Hochedlinger, K. & Jaenisch, R. Monoclonal mice generated by nuclear transfer from mature B and T donor cells. Nature 415, 1035–1038 (2002).
Lanza, R. P. et al. Cloning of an endangered species (Bos gaurus) using interspecies nuclear transfer. Cloning 2, 79–90 (2000).
Loi, P. et al. Genetic rescue of an endangered mammal by cross-species nuclear transfer using post-mortem somatic cells. Nat. Biotechnol. 19, 962–964 (2001).
Wakayama, S. et al. Propagation of an infertile hermaphrodite mouse lacking germ cells by using nuclear transfer and embryonic stem cell technology. Proc. Natl. Acad. Sci. USA 102, 29–33 (2005).
Vajta, G. & Gjerris, M. Science and technology of farm animal cloning: state of the art. Anim. Reprod. Sci. 92, 211–230 (2006).
Yang, X. et al. Risk assessment of meat and milk from cloned animals. Nat. Biotechnol. 25, 77–83 (2007).
Kato, Y. et al. Eight calves cloned from somatic cells of a single adult. Science 282, 2095–2098 (1998).
Hoshino, Y. et al. Resurrection of a bull by cloning from organs frozen without cryoprotectant in a −80 degrees c freezer for a decade. PLoS One 4, e4142 (2009).
Malin, K., Witkowska-Pilaszewicz, O. & Papis, K. The many problems of somatic cell nuclear transfer in reproductive cloning of mammals. Theriogenology 189, 246–254 (2022).
Matoba, S. & Zhang, Y. Somatic cell nuclear transfer reprogramming: mechanisms and applications. Cell Stem Cell 23, 471–485 (2018).
Yin, X. J. et al. Production of second-generation cloned cats by somatic cell nuclear transfer. Theriogenology 69, 1001–1006 (2008).
Wakayama, T. et al. Cloning of mice to six generations. Nature 407, 318–319 (2000).
Kubota, C., Tian, X. C. & Yang, X. Serial bull cloning by somatic cell nuclear transfer. Nat. Biotechnol. 22, 693–694 (2004).
Cho, S. K. et al. Serial cloning of pigs by somatic cell nuclear transfer: restoration of phenotypic normality during serial cloning. Dev. Dyn. 236, 3369–3382 (2007).
Kurome, M. et al. Production efficiency and telomere length of the cloned pigs following serial somatic cell nuclear transfer. J. Reprod. Dev. 54, 254–258 (2008).
Yang, M. et al. Increased pregnancy losses following serial somatic cell nuclear transfer in goats. Reprod. Fertil. Dev. 30, 1443–1453 (2018).
Kishigami, S. et al. Significant improvement of mouse cloning technique by treatment with trichostatin A after somatic nuclear transfer. Biochem. Biophys. Res Commun. 340, 183–189 (2006).
Wakayama, S. et al. Successful serial recloning in the mouse over multiple generations. Cell Stem Cell 12, 293–297 (2013).
Wakayama, T. & Yanagimachi, R. Cloning of male mice from adult tail-tip cells. Nat. Genet 22, 127–128 (1999).
Inoue, K. et al. Loss of H3K27me3 imprinting in the Sfmbt2 miRNA cluster causes enlargement of cloned mouse placentas. Nat. Commun. 11, 2150 (2020).
Tanaka, S. et al. Placentomegaly in cloned mouse concepti caused by expansion of the spongiotrophoblast layer. Biol. Reprod. 65, 1813–1821 (2001).
Wang, F., Kou, Z., Zhang, Y. & Gao, S. Dynamic reprogramming of histone acetylation and methylation in the first cell cycle of cloned mouse embryos. Biol. Reprod. 77, 1007–1016 (2007).
Terashita, Y. et al. Latrunculin a treatment prevents abnormal chromosome segregation for successful development of cloned embryos. PloS One 8, e78380 (2013).
Uchimura, A. et al. Early embryonic mutations reveal dynamics of somatic and germ cell lineages in mice. Genome Res. 32, 945–955 (2022).
Matsuda, Y. et al. Spectra and characteristics of somatic mutations induced by ionizing radiation in hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 120, e2216550120 (2023).
Sugo, N. et al. DNA polymerase beta suppresses somatic indels at CpG dinucleotides in developing cortical neurons. Proc. Natl. Acad. Sci. USA 122, e2506846122 (2025).
Porubsky, D. et al. Human de novo mutation rates from a four-generation pedigree reference. Nature 643, 427–436 (2025).
Belyeu, J. R. et al. De novo structural mutation rates and gamete-of-origin biases revealed through genome sequencing of 2,396 families. Am. J. Hum. Genet. 108, 597–607 (2021).
Inoue, K. et al. Trichostatin A specifically improves the aberrant expression of transcription factor genes in embryos produced by somatic cell nuclear transfer. Sci. Rep. 5, 10127 (2015).
Bouyahya, A. et al. Pharmacological properties of trichostatin a, focusing on the anticancer potential: a comprehensive review. Pharmaceuticals 15, 1235 (2022).
Mueller, T., Hantsch, C., Volkmer, I. & Staege, M. S. Differentiation-dependent regulation of human endogenous retrovirus K sequences and neighboring genes in germ cell tumor cells. Front. Microbiol. 9, 1253 (2018).
Dudko, N. et al. Oocyte age-dependent DNA damage can be reverted by the DNA repair competent karyoplasm of young oocytes. Aging Cell 25, e70300 (2025).
Valen, L. V. A new evolutionary law. Evol. Theory 1, 1–30 (1973).
Muller, H. J. The relation of recombination to mutational advance. Mutat. Res. 106, 2–9 (1964).
Michael, L. & Wilfried, G. Mutation load and the survival of small populations. Evolution 44, 1725–1737 (1990).
Wakayama, S. et al. Production of healthy cloned mice from bodies frozen at −20 degrees C for 16 years. Proc. Natl. Acad. Sci. USA 105, 17318–17322 (2008).
Mizutani, E. et al. Generation of cloned mice and nuclear transfer embryonic stem cell lines from urine-derived cells. Sci. Rep. 6, 23808 (2016).
Wakayama, S. & Wakayama, T. Can humanity thrive beyond the galaxy? J. Reprod. Dev. 71, 10–16 (2025).
Kimura, Y. & Yanagimachi, R. Intracytoplasmic sperm injection in the mouse. Biol. Reprod. 52, 709–720 (1995).
Chatot, C. L., Ziomek, C. A., Bavister, B. D., Lewis, J. L. & Torres, I. An improved culture medium supports development of random-bred 1-cell mouse embryos in vitro. J. Reprod. Fertil. 86, 679–688 (1989).
Wakayama, S., Ito, D., Hayashi, E., Ishiuchi, T. & Wakayama, T. Healthy cloned offspring derived from freeze-dried somatic cells. Nat. Commun. 13, 3666 (2022).
Ono, T. et al. Inhibition of class IIb histone deacetylase significantly improves cloning efficiency in mice. Biol. Reprod. 83, 929–937 (2010).
Van Thuan, N. et al. The histone deacetylase inhibitor scriptaid enhances nascent mRNA production and rescues full-term development in cloned inbred mice. Reprod. Camb. Engl. 138, 309–317 (2009).
Terashita, Y. et al. Latrunculin a can improve the birth rate of cloned mice and simplify the nuclear transfer protocol by gently inhibiting actin polymerization. Biol. Reprod. 86, 180 (2012).
Yang, L. L. et al. A novel, simplified method to prepare and preserve freeze-dried mouse sperm in plastic microtubes. J. Reprod. Dev. 69, 198–205 (2023).
Wakayama, S. et al. Healthy offspring from freeze-dried mouse spermatozoa held on the International Space Station for 9 months. Proc. Natl. Acad. Sci. USA 114, 5988–5993 (2017).
Hikichi, T. et al. Differentiation potential of parthenogenetic embryonic stem cells is improved by nuclear transfer. Stem Cells 25, 46–53 (2007).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Ostrander, B. E. P. et al. Whole-genome analysis for effective clinical diagnosis and gene discovery in early infantile epileptic encephalopathy. NPJ Genom. Med. 3, 22 (2018).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Jeffares, D. C. et al. Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat. Commun. 8, 14061 (2017).
Diaz-Gay, M. et al. Assigning mutational signatures to individual samples and individual somatic mutations with SigProfilerAssignment. Bioinformatics 39, btad756 (2023).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. Austin 6, 80–92 (2012).
Davydov, E. V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput. Biol. 6, e1001025 (2010).
Farmery, J. H. R., Smith, M. L., Diseases, N. B.-R. & Lynch, A. G. Telomerecat: a ploidy-agnostic method for estimating telomere length from whole genome sequencing data. Sci. Rep. 8, 1300 (2018).
Acknowledgements
We thank Dr. S. Kishigami, Mrs. Y. Kanda for assistance in preparing this manuscript. This work was partially funded by the Research Fellowships of Japan Society for the Promotion of Science to D.I. (JP20J23364), A.U. (24K03096, 24H02061, 24H00752), S. W. (23K08843) to T.W. (23K18124 and 24K01779); the Naito Foundation and Takahashi Industrial and Economic Research Foundation (189) to S.W.; Asada Science Foundation and the Canon Foundation (M20-0008) to T.W.
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Atsuo Ogura, Jie Xu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wakayama, S., Ito, D., Inoue, R. et al. Limitations of serial cloning in mammals. Nat Commun 17, 2495 (2026). https://doi.org/10.1038/s41467-026-69765-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69765-7